Probe CUST_19393_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19393_PI426222305 | JHI_St_60k_v1 | DMT400072760 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
All Microarray Probes Designed to Gene DMG400028315
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19183_PI426222305 | JHI_St_60k_v1 | DMT400072761 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
| CUST_19393_PI426222305 | JHI_St_60k_v1 | DMT400072760 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
Microarray Signals from CUST_19393_PI426222305
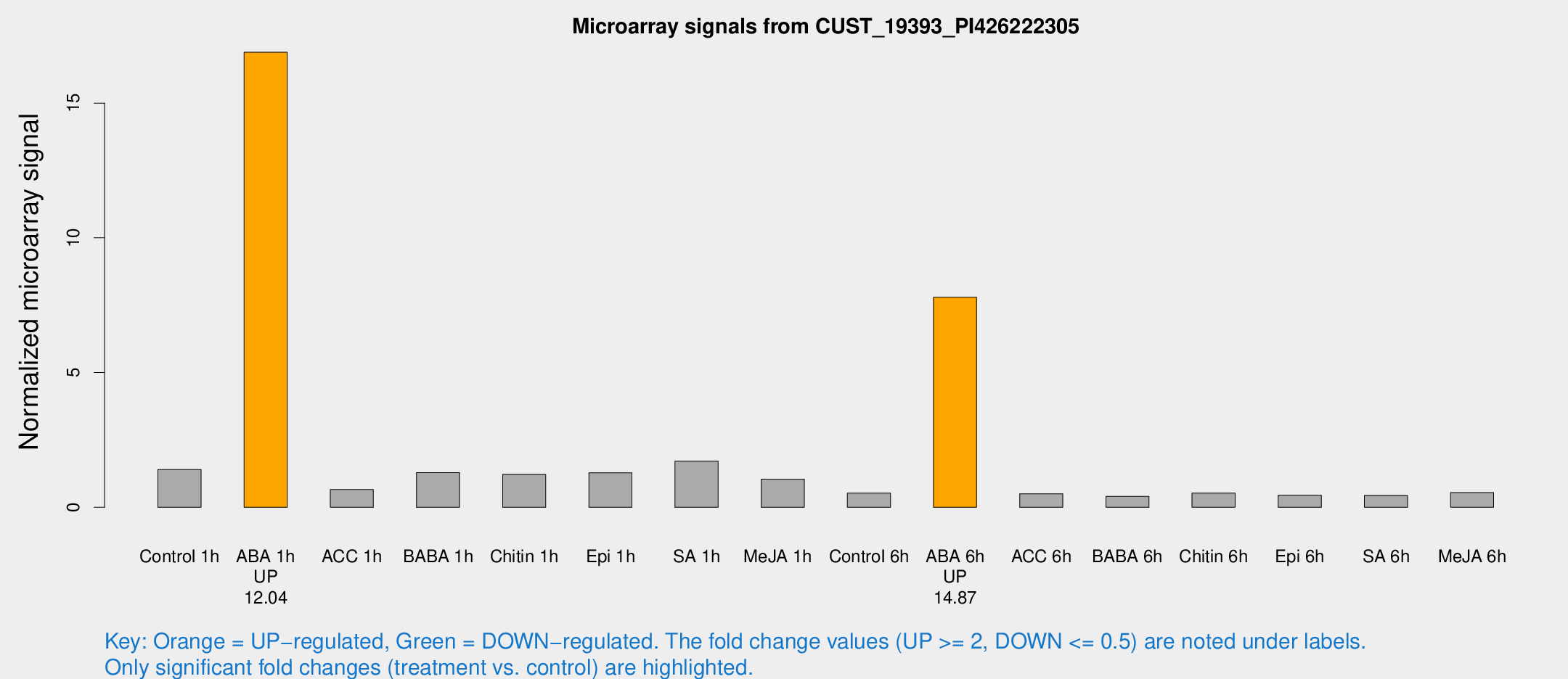
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 100.752 | 13.6341 | 1.40216 | 0.253349 |
| ABA 1h | 1070.76 | 131.596 | 16.8884 | 1.6134 |
| ACC 1h | 62.4553 | 23.8916 | 0.660512 | 0.416179 |
| BABA 1h | 92.3081 | 19.2287 | 1.28912 | 0.174423 |
| Chitin 1h | 78.0371 | 6.12709 | 1.22331 | 0.197047 |
| Epi 1h | 78.6069 | 5.95903 | 1.28052 | 0.0925545 |
| SA 1h | 124.66 | 7.97234 | 1.71486 | 0.109468 |
| Me-JA 1h | 60.3342 | 4.92631 | 1.04726 | 0.11553 |
| Control 6h | 37.2316 | 4.26384 | 0.524293 | 0.0609553 |
| ABA 6h | 601.037 | 94.7038 | 7.79547 | 0.980124 |
| ACC 6h | 41.1921 | 4.94253 | 0.503445 | 0.129025 |
| BABA 6h | 32.7574 | 4.90479 | 0.404544 | 0.0822101 |
| Chitin 6h | 39.9605 | 4.63162 | 0.52873 | 0.0619868 |
| Epi 6h | 38.1727 | 8.56483 | 0.455655 | 0.131663 |
| SA 6h | 32.6432 | 7.18786 | 0.440432 | 0.0678197 |
| Me-JA 6h | 47.105 | 22.307 | 0.546044 | 0.227789 |
Source Transcript PGSC0003DMT400072760 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G13740.1 | +2 | 5e-54 | 191 | 150/415 (36%) | ABI five binding protein 2 | chr1:4713969-4715158 FORWARD LENGTH=348 |
