Probe CUST_19340_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19340_PI426222305 | JHI_St_60k_v1 | DMT400072922 | CATGTCACCATCATCATCCATTAGATGTTCTATTTCATCACATACCTGAACATTTTTCGA |
All Microarray Probes Designed to Gene DMG400028369
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19340_PI426222305 | JHI_St_60k_v1 | DMT400072922 | CATGTCACCATCATCATCCATTAGATGTTCTATTTCATCACATACCTGAACATTTTTCGA |
| CUST_19416_PI426222305 | JHI_St_60k_v1 | DMT400072921 | TAAGCAGCCTACCTTGATGCAATTGTTAAAGACAGAATCCATGGCTCAAACCCGTGAGCT |
Microarray Signals from CUST_19340_PI426222305
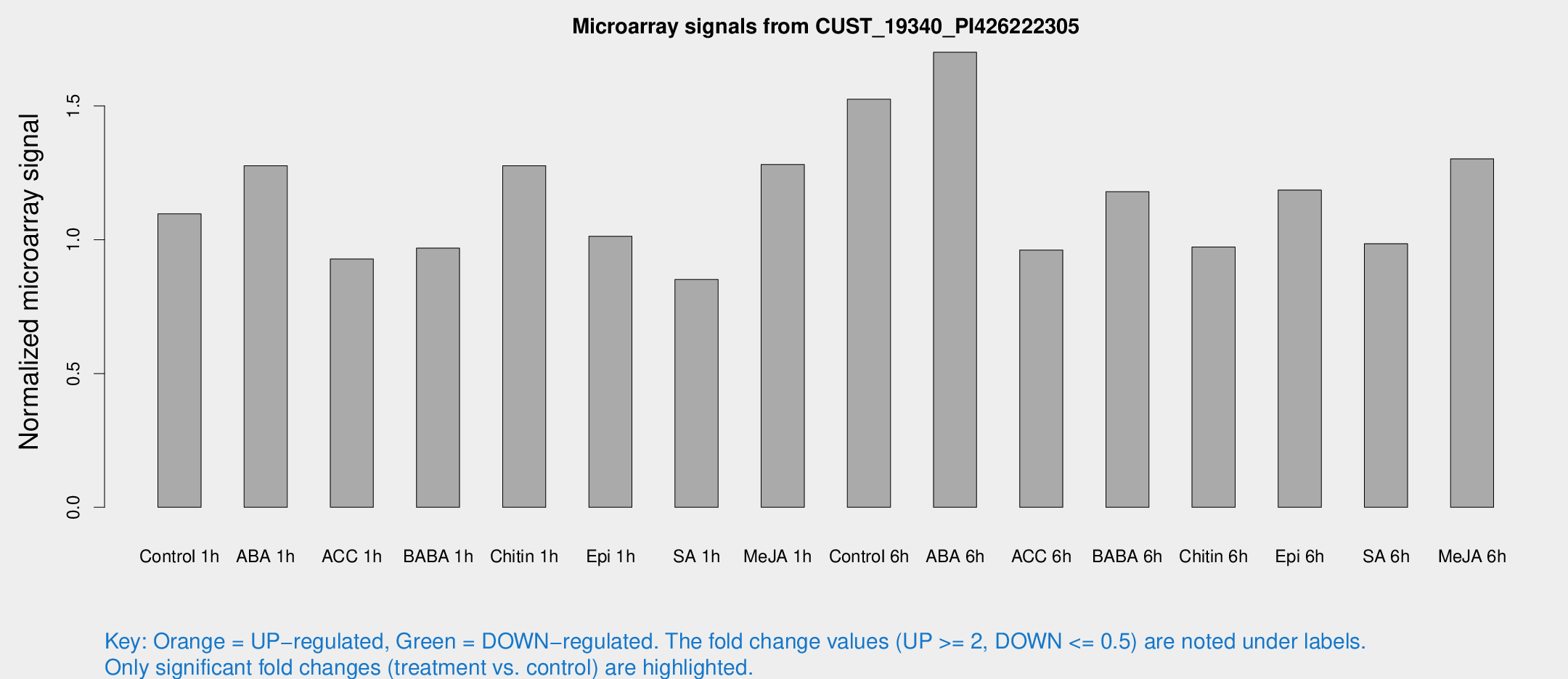
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.76574 | 3.36845 | 1.09683 | 0.53493 |
| ABA 1h | 8.04732 | 3.32435 | 1.27632 | 0.611309 |
| ACC 1h | 6.38354 | 3.71008 | 0.928159 | 0.538335 |
| BABA 1h | 6.24118 | 3.62232 | 0.968643 | 0.562049 |
| Chitin 1h | 8.00531 | 3.48378 | 1.27674 | 0.6184 |
| Epi 1h | 5.86999 | 3.40683 | 1.01309 | 0.586967 |
| SA 1h | 5.85081 | 3.39029 | 0.851861 | 0.493375 |
| Me-JA 1h | 7.27011 | 3.50998 | 1.28115 | 0.655787 |
| Control 6h | 13.1409 | 6.83391 | 1.5251 | 0.76194 |
| ABA 6h | 12.3613 | 3.80187 | 1.70091 | 0.547345 |
| ACC 6h | 7.54088 | 4.4018 | 0.961524 | 0.544822 |
| BABA 6h | 9.36798 | 4.06645 | 1.17974 | 0.57526 |
| Chitin 6h | 6.92427 | 3.9334 | 0.972848 | 0.553093 |
| Epi 6h | 9.31927 | 4.18638 | 1.18602 | 0.555946 |
| SA 6h | 6.51535 | 3.77983 | 0.985199 | 0.571309 |
| Me-JA 6h | 9.89207 | 3.77983 | 1.30251 | 0.613568 |
Source Transcript PGSC0003DMT400072922 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G03620.1 | -3 | 9e-24 | 105 | 51/75 (68%) | magnesium transporter 3 | chr2:1100489-1102168 REVERSE LENGTH=421 |
