Probe CUST_19223_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19223_PI426222305 | JHI_St_60k_v1 | DMT400072742 | GCTGTGGCCTTTACATCTCGAAGATTAATGACATAATGCTAGTGTTTTTTCGAGATAATA |
All Microarray Probes Designed to Gene DMG400028306
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19335_PI426222305 | JHI_St_60k_v1 | DMT400072741 | CCCACATGCCTCGAAGGTAGCTAACATTATTTTACAATTTTGTGATTGTAAAGAGCTATT |
| CUST_19223_PI426222305 | JHI_St_60k_v1 | DMT400072742 | GCTGTGGCCTTTACATCTCGAAGATTAATGACATAATGCTAGTGTTTTTTCGAGATAATA |
Microarray Signals from CUST_19223_PI426222305
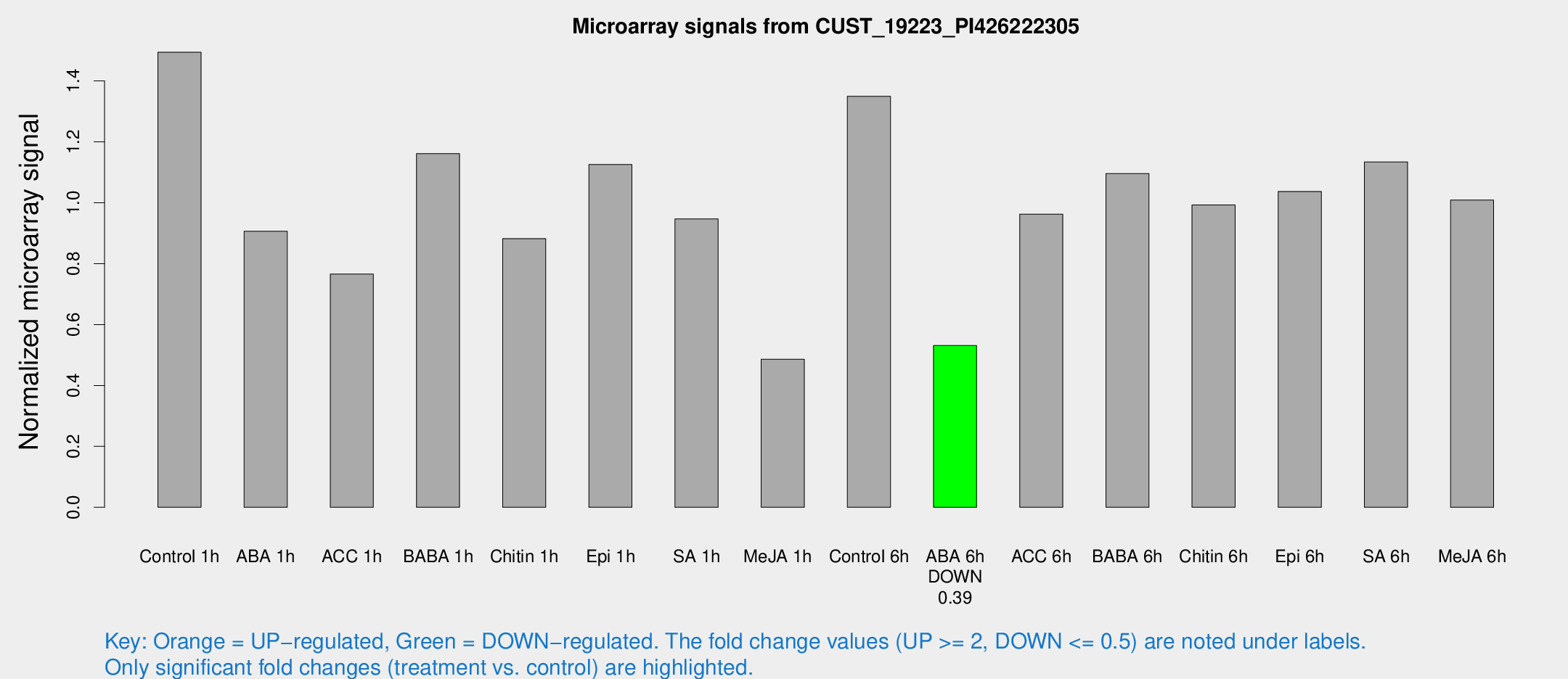
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 106.123 | 14.3768 | 1.49437 | 0.129666 |
| ABA 1h | 56.4001 | 4.85583 | 0.906524 | 0.0763373 |
| ACC 1h | 59.1086 | 14.1452 | 0.765949 | 0.16569 |
| BABA 1h | 82.1769 | 17.5814 | 1.16129 | 0.158571 |
| Chitin 1h | 56.2396 | 6.71458 | 0.882372 | 0.0783456 |
| Epi 1h | 71.8183 | 15.132 | 1.12547 | 0.270616 |
| SA 1h | 69.1327 | 9.14261 | 0.946972 | 0.106048 |
| Me-JA 1h | 30.4449 | 8.69881 | 0.486196 | 0.117888 |
| Control 6h | 96.6099 | 14.453 | 1.34976 | 0.128351 |
| ABA 6h | 39.444 | 4.40403 | 0.531271 | 0.0595011 |
| ACC 6h | 82.2461 | 18.584 | 0.962855 | 0.20846 |
| BABA 6h | 86.1567 | 6.59541 | 1.09586 | 0.0818731 |
| Chitin 6h | 75.7621 | 12.0253 | 0.992811 | 0.156594 |
| Epi 6h | 83.0707 | 10.5294 | 1.03728 | 0.0801481 |
| SA 6h | 80.2891 | 13.2857 | 1.1338 | 0.154199 |
| Me-JA 6h | 70.9228 | 7.55783 | 1.00886 | 0.0935533 |
Source Transcript PGSC0003DMT400072742 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G02290.1 | +3 | 5e-38 | 140 | 96/298 (32%) | RING/U-box superfamily protein | chr3:459385-460688 FORWARD LENGTH=231 |
