Probe CUST_19183_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19183_PI426222305 | JHI_St_60k_v1 | DMT400072761 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
All Microarray Probes Designed to Gene DMG400028315
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19183_PI426222305 | JHI_St_60k_v1 | DMT400072761 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
| CUST_19393_PI426222305 | JHI_St_60k_v1 | DMT400072760 | TCTTGGGACCATATTGTATGATTGCATCATTGTGTTGTTGGGGGAGACAAAGAAGTGTTT |
Microarray Signals from CUST_19183_PI426222305
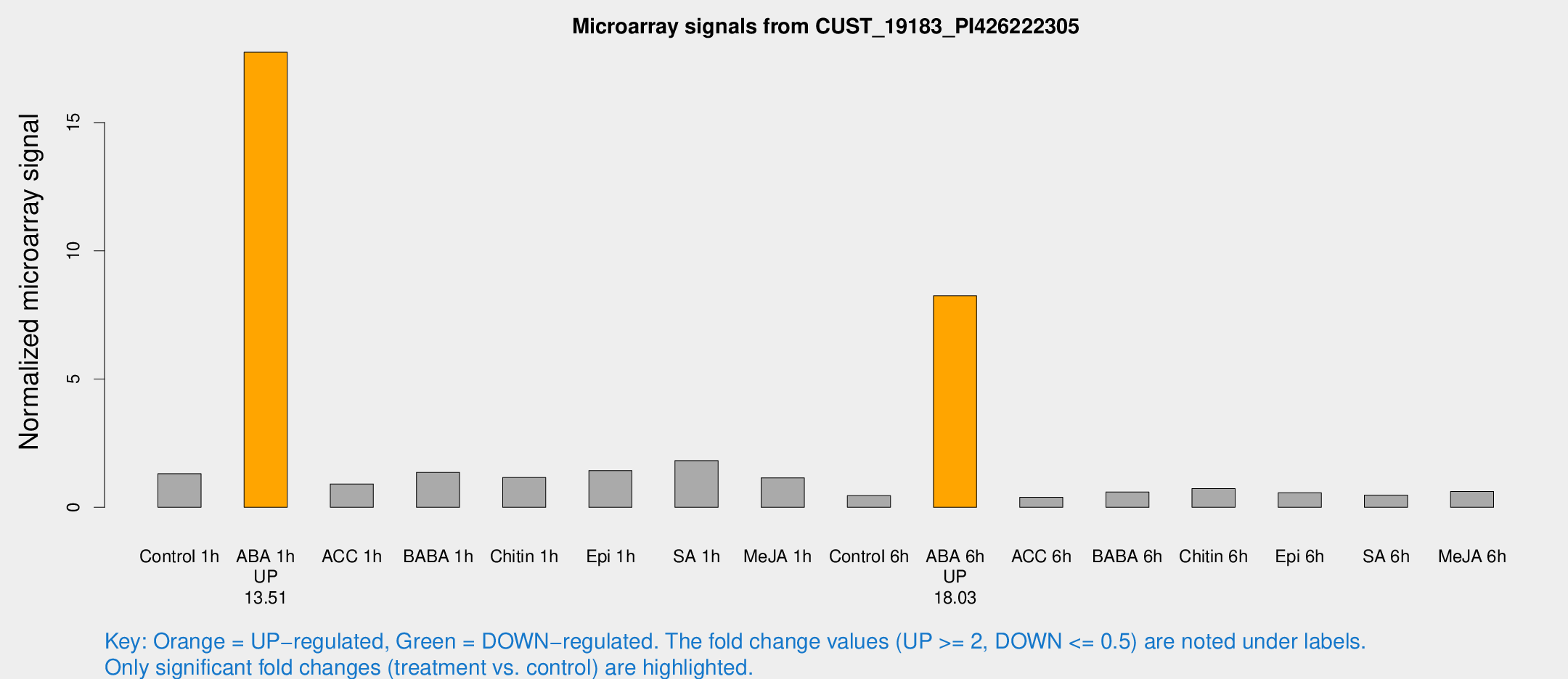
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 93.2601 | 10.8138 | 1.3131 | 0.172172 |
| ABA 1h | 1118.74 | 144.981 | 17.742 | 1.83501 |
| ACC 1h | 71.0923 | 17.7956 | 0.91073 | 0.2143 |
| BABA 1h | 92.7842 | 8.72413 | 1.3629 | 0.0976512 |
| Chitin 1h | 74.0734 | 8.36129 | 1.16088 | 0.222725 |
| Epi 1h | 88.6294 | 12.2447 | 1.43256 | 0.20063 |
| SA 1h | 133.25 | 18.1913 | 1.82126 | 0.253226 |
| Me-JA 1h | 66.4503 | 7.96324 | 1.14596 | 0.209147 |
| Control 6h | 32.5171 | 4.26472 | 0.457274 | 0.0623855 |
| ABA 6h | 623.596 | 76.7792 | 8.24365 | 0.83102 |
| ACC 6h | 33.5198 | 7.91396 | 0.390155 | 0.178888 |
| BABA 6h | 46.9764 | 4.99028 | 0.595806 | 0.0632425 |
| Chitin 6h | 54.3797 | 5.23377 | 0.728346 | 0.0703737 |
| Epi 6h | 47.087 | 10.9031 | 0.568067 | 0.0887421 |
| SA 6h | 38.9784 | 12.8606 | 0.476223 | 0.171498 |
| Me-JA 6h | 48.2931 | 16.4401 | 0.616838 | 0.204781 |
Source Transcript PGSC0003DMT400072761 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G29575.1 | +3 | 7e-34 | 133 | 64/94 (68%) | ABI five binding protein 3 | chr3:11382416-11383657 REVERSE LENGTH=231 |
