Probe CUST_19042_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19042_PI426222305 | JHI_St_60k_v1 | DMT400052621 | GCAGAAGAACATGGAAATACTCAAATTTCAACTGTCATTTTTGATTTGGATGGTACCCTT |
All Microarray Probes Designed to Gene DMG400020426
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19042_PI426222305 | JHI_St_60k_v1 | DMT400052621 | GCAGAAGAACATGGAAATACTCAAATTTCAACTGTCATTTTTGATTTGGATGGTACCCTT |
| CUST_19050_PI426222305 | JHI_St_60k_v1 | DMT400052620 | GTTATAAAGTGTTTATTGCAGCATTACTCAATGTGTCTGTGCAGTGGACAACTGGTATCT |
Microarray Signals from CUST_19042_PI426222305
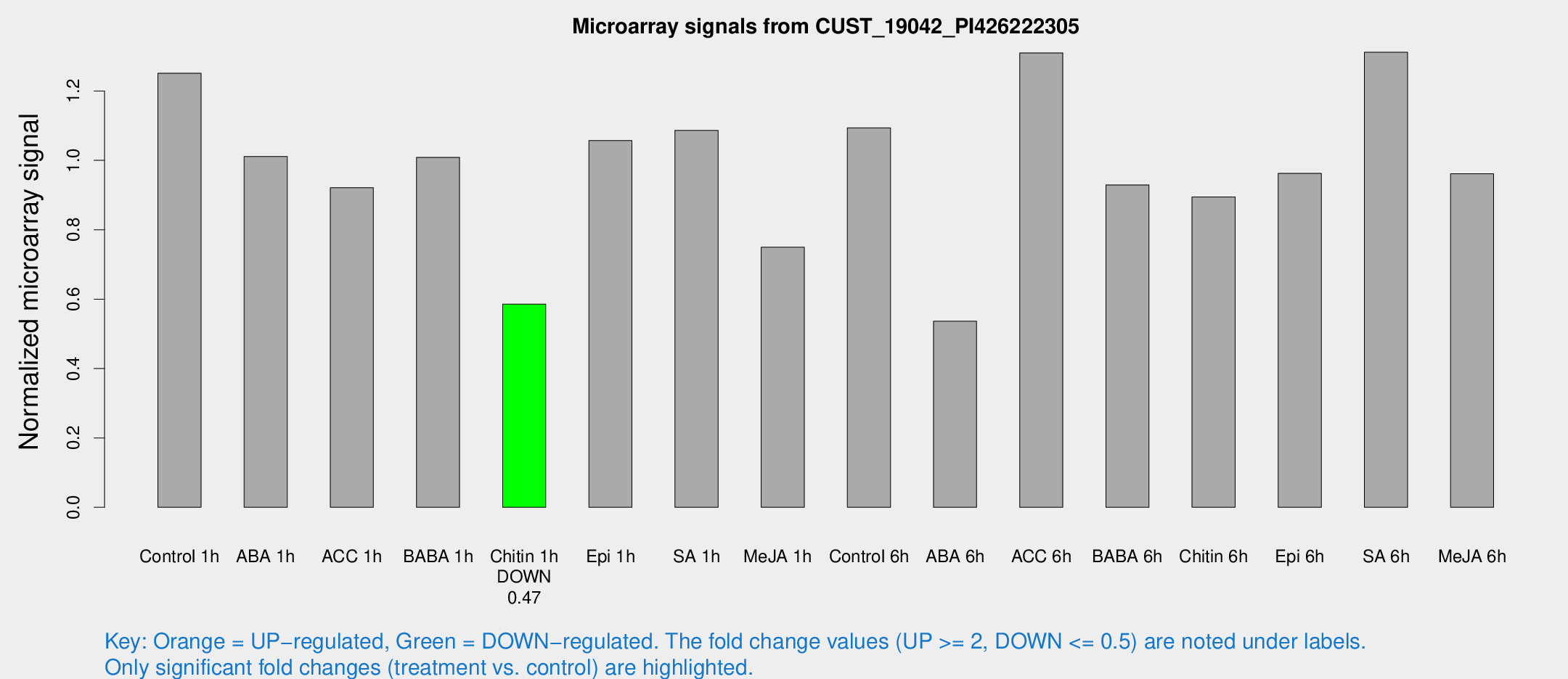
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 60.0846 | 9.60922 | 1.25121 | 0.114045 |
| ABA 1h | 42.3767 | 4.59114 | 1.0114 | 0.0965702 |
| ACC 1h | 49.5612 | 14.7642 | 0.921335 | 0.274682 |
| BABA 1h | 49.1808 | 12.7384 | 1.00876 | 0.197646 |
| Chitin 1h | 24.741 | 3.61607 | 0.585443 | 0.0849849 |
| Epi 1h | 44.3497 | 7.81759 | 1.05728 | 0.180378 |
| SA 1h | 55.3064 | 11.9964 | 1.08622 | 0.211514 |
| Me-JA 1h | 29.0199 | 3.76394 | 0.749781 | 0.0978817 |
| Control 6h | 57.1489 | 18.7469 | 1.09385 | 0.307317 |
| ABA 6h | 36.5148 | 14.6116 | 0.536428 | 0.435852 |
| ACC 6h | 71.903 | 10.2611 | 1.30984 | 0.231604 |
| BABA 6h | 52.9359 | 14.5546 | 0.929258 | 0.261293 |
| Chitin 6h | 47.6991 | 12.5168 | 0.894685 | 0.22118 |
| Epi 6h | 56.1207 | 18.1683 | 0.962699 | 0.344984 |
| SA 6h | 64.6758 | 16.7208 | 1.31176 | 0.526656 |
| Me-JA 6h | 48.6385 | 12.1143 | 0.961637 | 0.223045 |
Source Transcript PGSC0003DMT400052621 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G21470.1 | +3 | 7e-81 | 261 | 159/362 (44%) | riboflavin kinase/FMN hydrolase | chr4:11431284-11433197 FORWARD LENGTH=379 |
