Probe CUST_19021_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19021_PI426222305 | JHI_St_60k_v1 | DMT400044514 | CCCAGTATGGTGCTCCTCTTATATTGCATAGATTGTAAGCTATAGCATAATTAAGTTGTA |
All Microarray Probes Designed to Gene DMG400017281
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19021_PI426222305 | JHI_St_60k_v1 | DMT400044514 | CCCAGTATGGTGCTCCTCTTATATTGCATAGATTGTAAGCTATAGCATAATTAAGTTGTA |
Microarray Signals from CUST_19021_PI426222305
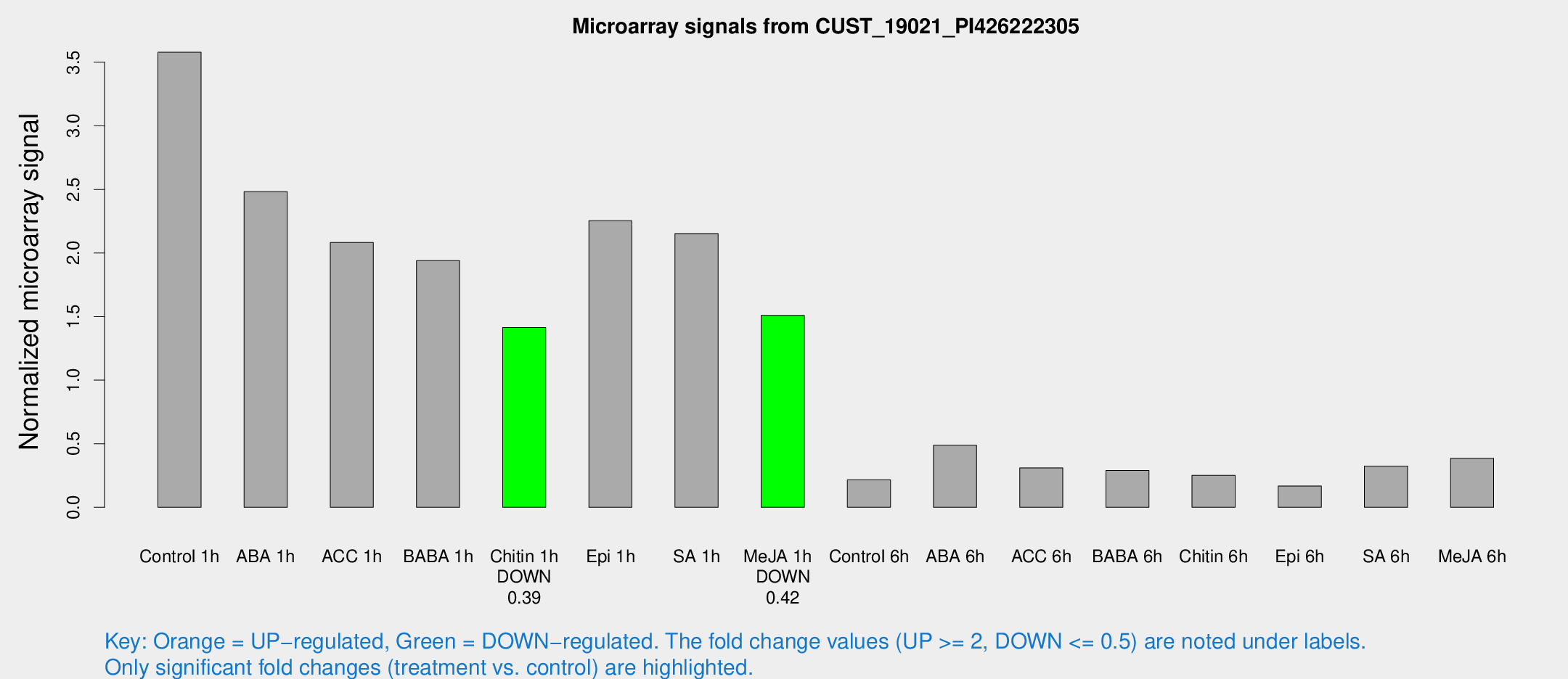
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5493.4 | 661.856 | 3.57899 | 0.206643 |
| ABA 1h | 3414.2 | 539.229 | 2.48266 | 0.3391 |
| ACC 1h | 3484.18 | 918.004 | 2.08218 | 0.463388 |
| BABA 1h | 3097.34 | 805.475 | 1.93998 | 0.431867 |
| Chitin 1h | 1956.08 | 263.174 | 1.41322 | 0.0816233 |
| Epi 1h | 2982.31 | 312.179 | 2.25335 | 0.276427 |
| SA 1h | 3422.62 | 485.836 | 2.15265 | 0.222248 |
| Me-JA 1h | 1890.14 | 218.217 | 1.50934 | 0.0871791 |
| Control 6h | 388.042 | 169.192 | 0.215258 | 0.0891652 |
| ABA 6h | 833.159 | 197.036 | 0.487648 | 0.114374 |
| ACC 6h | 540.042 | 31.4757 | 0.310474 | 0.0285375 |
| BABA 6h | 495.191 | 38.7847 | 0.290662 | 0.0196939 |
| Chitin 6h | 439.364 | 129.564 | 0.251146 | 0.0693662 |
| Epi 6h | 308.515 | 87.6938 | 0.167617 | 0.0345954 |
| SA 6h | 520.639 | 140.512 | 0.32367 | 0.0599811 |
| Me-JA 6h | 601.211 | 112.053 | 0.384953 | 0.100795 |
Source Transcript PGSC0003DMT400044514 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14690.1 | +2 | 0.0 | 604 | 290/512 (57%) | cytochrome P450, family 72, subfamily A, polypeptide 15 | chr3:4937410-4939310 FORWARD LENGTH=512 |
