Probe CUST_18590_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18590_PI426222305 | JHI_St_60k_v1 | DMT400050753 | CTGATTTTCATCGAGCTATCACCTATGGCTTCGAGCTTTATTATATCGGGTTTATAGAAA |
All Microarray Probes Designed to Gene DMG400019712
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18435_PI426222305 | JHI_St_60k_v1 | DMT400050751 | TTTTCTTCTGATTCAATGGCTTTGTTAGAAAATTCTGCTGATAATCCCATCGAAGTAGTC |
| CUST_18590_PI426222305 | JHI_St_60k_v1 | DMT400050753 | CTGATTTTCATCGAGCTATCACCTATGGCTTCGAGCTTTATTATATCGGGTTTATAGAAA |
Microarray Signals from CUST_18590_PI426222305
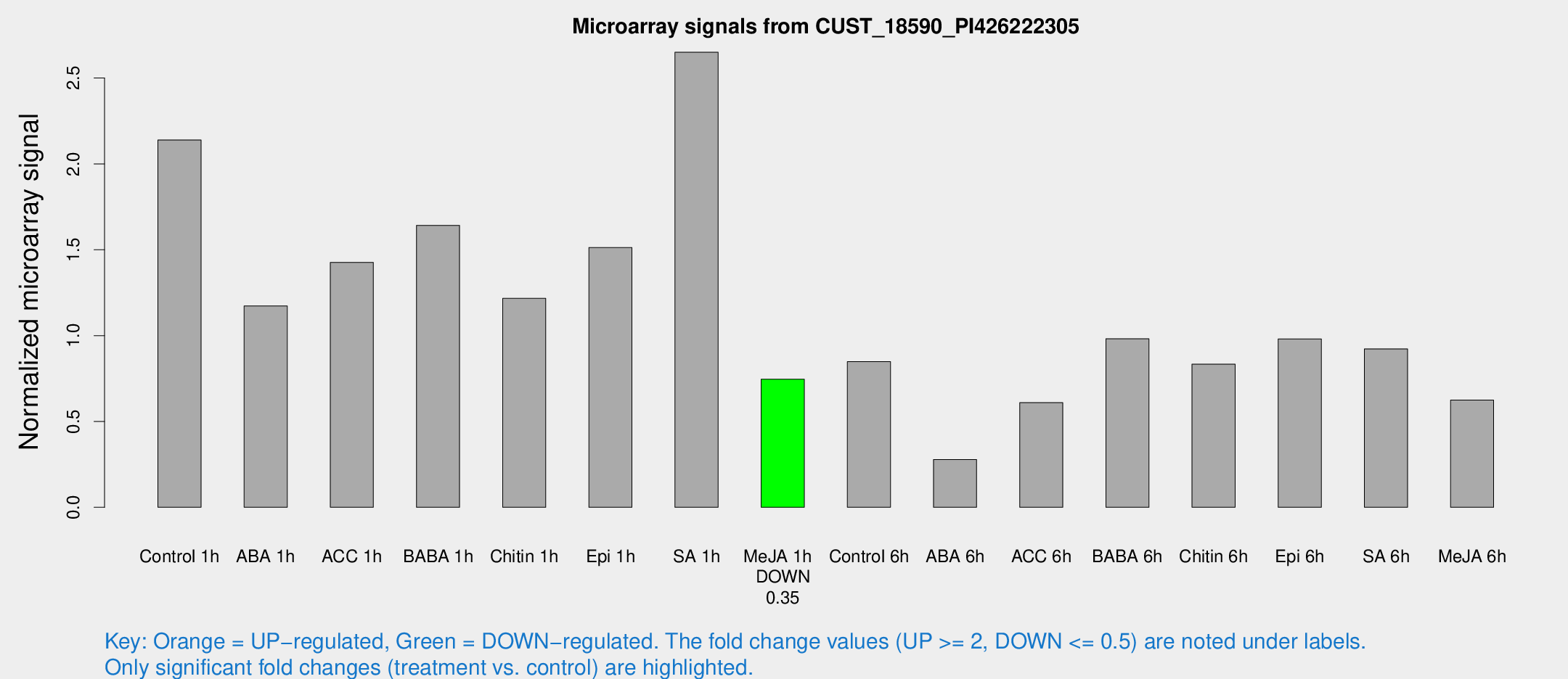
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 654.565 | 111.471 | 2.13968 | 0.215374 |
| ABA 1h | 325.699 | 74.9863 | 1.17307 | 0.190828 |
| ACC 1h | 502.639 | 155.605 | 1.42664 | 0.514989 |
| BABA 1h | 497.345 | 105.482 | 1.64224 | 0.229379 |
| Chitin 1h | 328.837 | 30.6498 | 1.21712 | 0.134089 |
| Epi 1h | 392.597 | 32.5051 | 1.51284 | 0.0959942 |
| SA 1h | 840.215 | 165.437 | 2.6505 | 0.387039 |
| Me-JA 1h | 183.699 | 21.8315 | 0.7458 | 0.0457592 |
| Control 6h | 258.256 | 35.2323 | 0.848509 | 0.0621269 |
| ABA 6h | 91.6424 | 18.8285 | 0.278571 | 0.0407557 |
| ACC 6h | 209.708 | 16.8989 | 0.60978 | 0.0683551 |
| BABA 6h | 328.435 | 20.8924 | 0.981566 | 0.0581253 |
| Chitin 6h | 271.937 | 43.2108 | 0.833519 | 0.153897 |
| Epi 6h | 337.858 | 50.9865 | 0.980602 | 0.0972254 |
| SA 6h | 274.515 | 27.8823 | 0.922653 | 0.0584077 |
| Me-JA 6h | 222.482 | 98.4029 | 0.624069 | 0.233187 |
Source Transcript PGSC0003DMT400050753 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G66910.1 | +3 | 2e-50 | 143 | 69/123 (56%) | Protein kinase superfamily protein | chr1:24961634-24963941 REVERSE LENGTH=666 |
