Probe CUST_18435_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18435_PI426222305 | JHI_St_60k_v1 | DMT400050751 | TTTTCTTCTGATTCAATGGCTTTGTTAGAAAATTCTGCTGATAATCCCATCGAAGTAGTC |
All Microarray Probes Designed to Gene DMG400019712
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18435_PI426222305 | JHI_St_60k_v1 | DMT400050751 | TTTTCTTCTGATTCAATGGCTTTGTTAGAAAATTCTGCTGATAATCCCATCGAAGTAGTC |
| CUST_18590_PI426222305 | JHI_St_60k_v1 | DMT400050753 | CTGATTTTCATCGAGCTATCACCTATGGCTTCGAGCTTTATTATATCGGGTTTATAGAAA |
Microarray Signals from CUST_18435_PI426222305
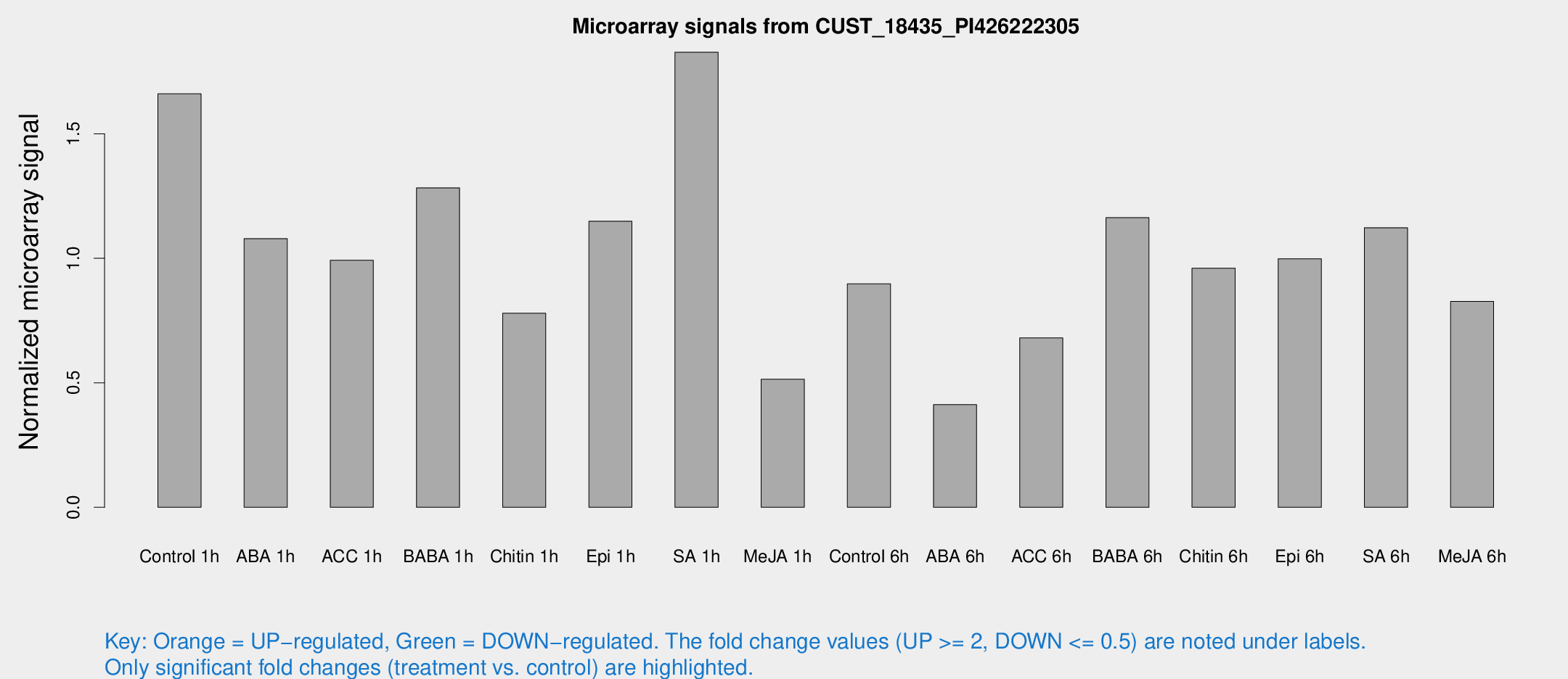
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2747.26 | 663.46 | 1.66053 | 0.280499 |
| ABA 1h | 1536.23 | 240.047 | 1.07898 | 0.116094 |
| ACC 1h | 2000.72 | 733.86 | 0.991851 | 0.534002 |
| BABA 1h | 2054.91 | 448.592 | 1.28283 | 0.194876 |
| Chitin 1h | 1132.76 | 206.833 | 0.779428 | 0.0783479 |
| Epi 1h | 1571.64 | 146.544 | 1.14886 | 0.0868067 |
| SA 1h | 3016.44 | 495.655 | 1.82756 | 0.183148 |
| Me-JA 1h | 690.739 | 159.344 | 0.514256 | 0.0714553 |
| Control 6h | 1487.07 | 312.587 | 0.897435 | 0.138358 |
| ABA 6h | 730.947 | 185.557 | 0.4124 | 0.0814121 |
| ACC 6h | 1230.14 | 97.4683 | 0.680193 | 0.092904 |
| BABA 6h | 2068.66 | 244.176 | 1.16306 | 0.0939757 |
| Chitin 6h | 1629.11 | 205.496 | 0.960468 | 0.124173 |
| Epi 6h | 1796.21 | 233.528 | 0.998136 | 0.0824985 |
| SA 6h | 1823.49 | 406.295 | 1.12275 | 0.165995 |
| Me-JA 6h | 1505.41 | 596.397 | 0.826931 | 0.274159 |
Source Transcript PGSC0003DMT400050751 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G38280.1 | +1 | 1e-13 | 67 | 33/79 (42%) | PR5-like receptor kinase | chr5:15293325-15295838 REVERSE LENGTH=665 |
