Probe CUST_18378_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18378_PI426222305 | JHI_St_60k_v1 | DMT400008935 | GTTGCCCTTTCTCTGTGAAAAGAAATCACTAATACTTTTTATGTGTCGATGTTGTGGATT |
All Microarray Probes Designed to Gene DMG402003479
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18378_PI426222305 | JHI_St_60k_v1 | DMT400008935 | GTTGCCCTTTCTCTGTGAAAAGAAATCACTAATACTTTTTATGTGTCGATGTTGTGGATT |
| CUST_18247_PI426222305 | JHI_St_60k_v1 | DMT400008936 | CCAAATTCCTATATTGTTTTATCAGCAGATGCTTTCAGGGCATGGACAAATGATAGTTTG |
Microarray Signals from CUST_18378_PI426222305
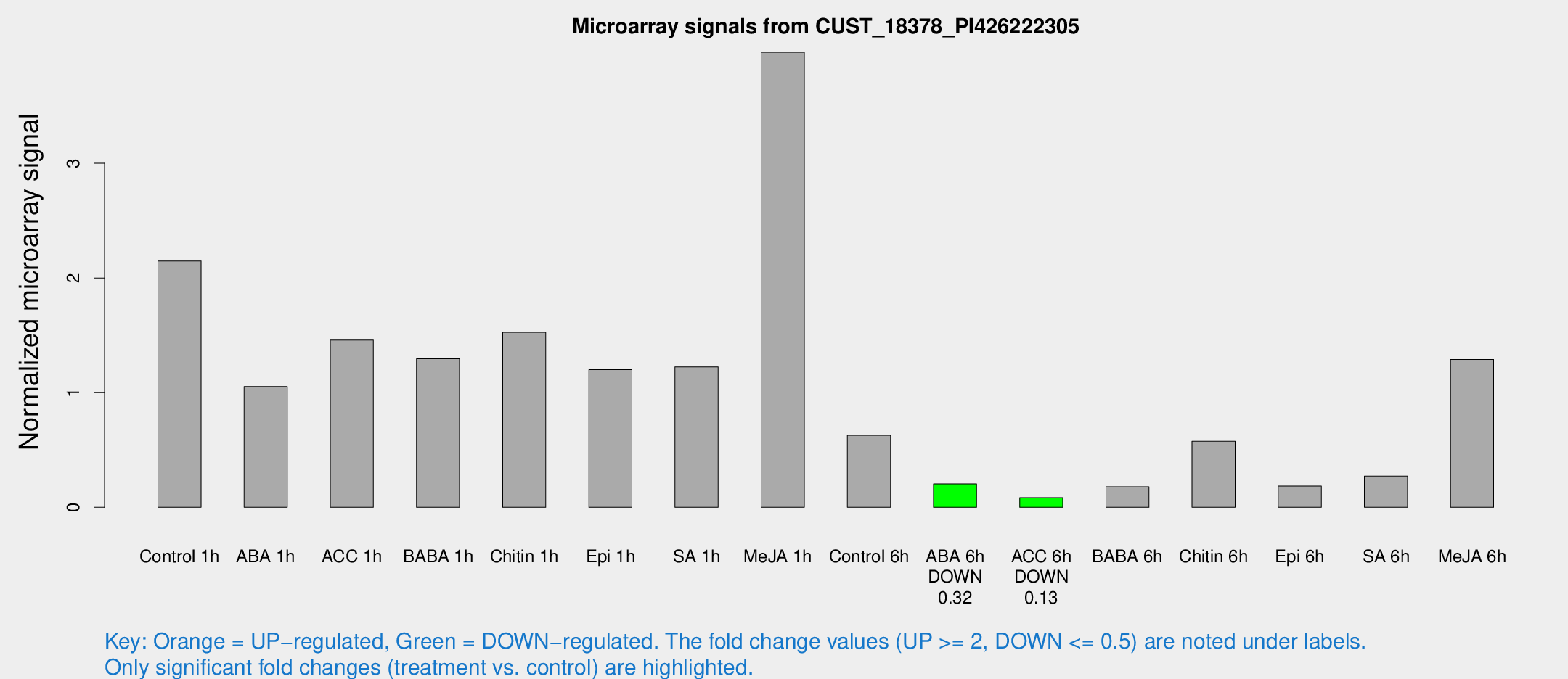
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 630.359 | 36.5834 | 2.14891 | 0.218101 |
| ABA 1h | 287.972 | 60.5432 | 1.05313 | 0.267911 |
| ACC 1h | 453.516 | 77.8994 | 1.45829 | 0.160355 |
| BABA 1h | 368.033 | 28.8795 | 1.29598 | 0.095562 |
| Chitin 1h | 412.843 | 70.6479 | 1.52587 | 0.290208 |
| Epi 1h | 311.564 | 48.6786 | 1.2002 | 0.211581 |
| SA 1h | 421.361 | 152.394 | 1.22464 | 0.551903 |
| Me-JA 1h | 976.796 | 168.59 | 3.96837 | 0.333204 |
| Control 6h | 214.895 | 82.6808 | 0.628176 | 0.208996 |
| ABA 6h | 65.4984 | 11.692 | 0.203772 | 0.0496931 |
| ACC 6h | 28.4205 | 4.28509 | 0.0837802 | 0.0126459 |
| BABA 6h | 116.388 | 57.0891 | 0.178872 | 0.602558 |
| Chitin 6h | 181.689 | 21.0423 | 0.575458 | 0.0823376 |
| Epi 6h | 118.993 | 58.7443 | 0.184763 | 0.660851 |
| SA 6h | 82.4852 | 18.4377 | 0.272041 | 0.0464411 |
| Me-JA 6h | 419.113 | 143.351 | 1.28956 | 0.3725 |
Source Transcript PGSC0003DMT400008935 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52820.1 | +3 | 1e-43 | 159 | 86/226 (38%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:19669216-19670321 FORWARD LENGTH=317 |
