Probe CUST_18352_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18352_PI426222305 | JHI_St_60k_v1 | DMT400055038 | AAACTTGCTCCTGATAAAGAGAAATCTGTGGTCACCAGGCTGGCCGGAACTTTTGGATAT |
All Microarray Probes Designed to Gene DMG400021353
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18352_PI426222305 | JHI_St_60k_v1 | DMT400055038 | AAACTTGCTCCTGATAAAGAGAAATCTGTGGTCACCAGGCTGGCCGGAACTTTTGGATAT |
| CUST_18229_PI426222305 | JHI_St_60k_v1 | DMT400055037 | ATGTTGTTGCAACAATGGTGTCACTTACACATCTTTGGCTTCATGGGAACCAATTTTCAG |
Microarray Signals from CUST_18352_PI426222305
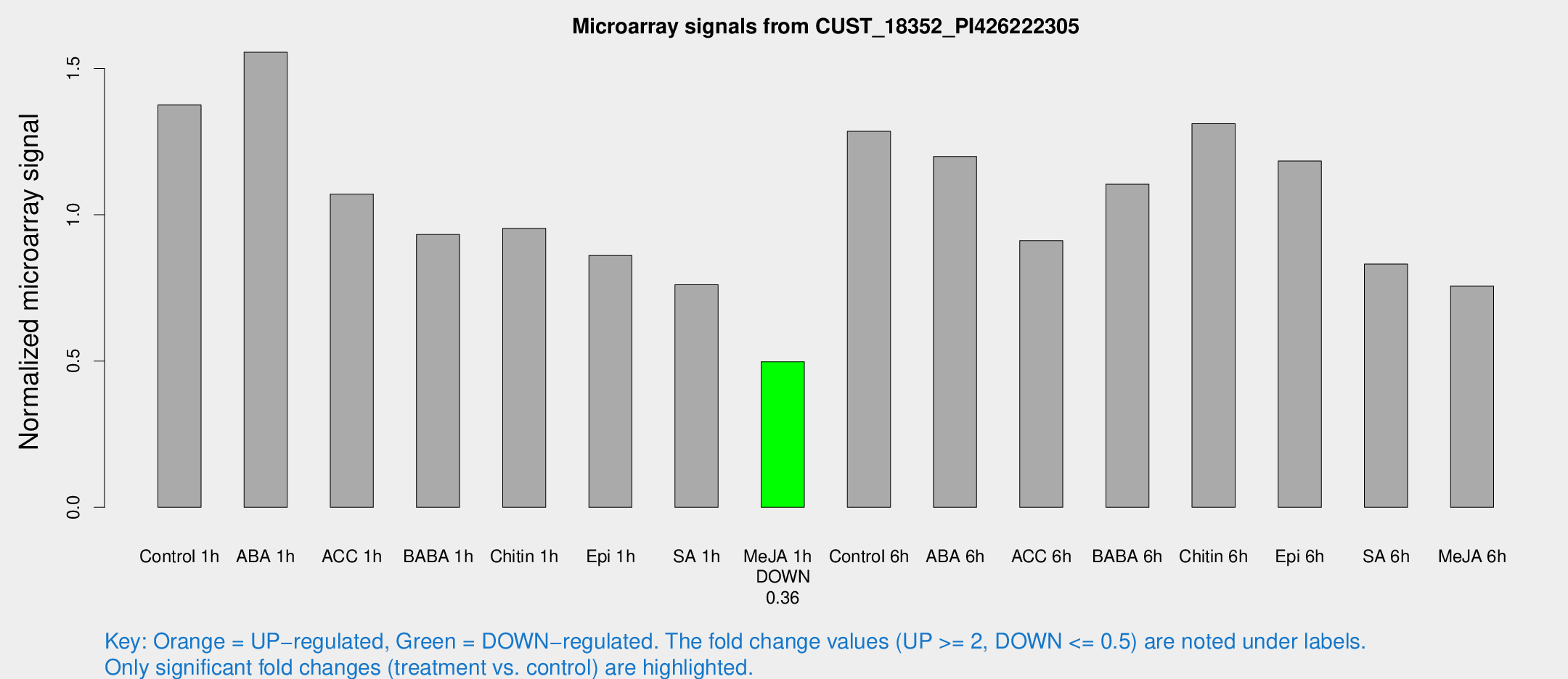
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 513.983 | 84.9919 | 1.37539 | 0.147506 |
| ABA 1h | 504.273 | 48.8523 | 1.55571 | 0.0904991 |
| ACC 1h | 409.344 | 66.656 | 1.07105 | 0.122319 |
| BABA 1h | 330.775 | 37.3788 | 0.9326 | 0.0549994 |
| Chitin 1h | 312.791 | 21.1303 | 0.953583 | 0.0615276 |
| Epi 1h | 271.998 | 19.3471 | 0.86093 | 0.051073 |
| SA 1h | 283.827 | 16.7929 | 0.760703 | 0.0535348 |
| Me-JA 1h | 147.758 | 9.34654 | 0.497719 | 0.0426085 |
| Control 6h | 499.631 | 116.208 | 1.28521 | 0.238375 |
| ABA 6h | 467.175 | 45.9572 | 1.1992 | 0.17566 |
| ACC 6h | 397.161 | 86.3932 | 0.911374 | 0.0719845 |
| BABA 6h | 455.329 | 55.1342 | 1.10453 | 0.115113 |
| Chitin 6h | 508.554 | 29.7666 | 1.3117 | 0.07656 |
| Epi 6h | 496.364 | 79.6281 | 1.18391 | 0.195733 |
| SA 6h | 330.412 | 90.1309 | 0.831423 | 0.190598 |
| Me-JA 6h | 306.719 | 87.5398 | 0.756729 | 0.231067 |
Source Transcript PGSC0003DMT400055038 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G66150.1 | +3 | 0.0 | 748 | 438/976 (45%) | transmembrane kinase 1 | chr1:24631503-24634415 FORWARD LENGTH=942 |
