Probe CUST_18302_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18302_PI426222305 | JHI_St_60k_v1 | DMT400042159 | AATGAGAAGGCACAAGAAAAGACATTAGTGTGTAACCTCGAGGTATCAACTTAATCTCTA |
All Microarray Probes Designed to Gene DMG400016357
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18302_PI426222305 | JHI_St_60k_v1 | DMT400042159 | AATGAGAAGGCACAAGAAAAGACATTAGTGTGTAACCTCGAGGTATCAACTTAATCTCTA |
| CUST_18274_PI426222305 | JHI_St_60k_v1 | DMT400042158 | ATTGGATCTCTCATCCAACAAAATCAGCGGAGAAATTCCACAACAACTTGCATCCCTCAA |
Microarray Signals from CUST_18302_PI426222305
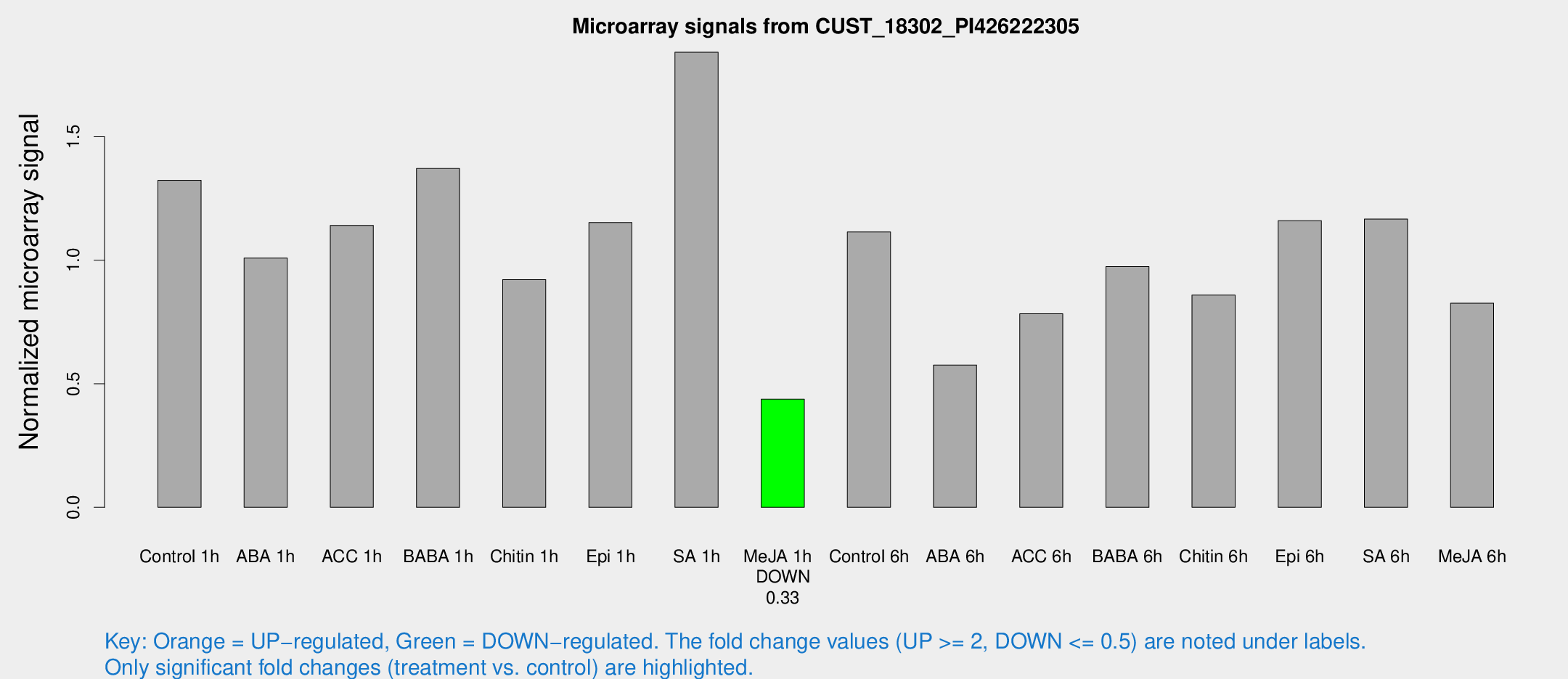
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1404.95 | 247.416 | 1.32355 | 0.168154 |
| ABA 1h | 926.771 | 76.3792 | 1.00879 | 0.0583456 |
| ACC 1h | 1336.34 | 364.608 | 1.14126 | 0.311981 |
| BABA 1h | 1436.91 | 299.035 | 1.37138 | 0.186959 |
| Chitin 1h | 865.033 | 98.2495 | 0.92173 | 0.0548843 |
| Epi 1h | 1036.55 | 87.7133 | 1.15268 | 0.0898674 |
| SA 1h | 1994.99 | 297.499 | 1.84219 | 0.160473 |
| Me-JA 1h | 381.361 | 73.394 | 0.437665 | 0.0453245 |
| Control 6h | 1176.48 | 161.977 | 1.11489 | 0.073669 |
| ABA 6h | 669.196 | 160.081 | 0.575825 | 0.110712 |
| ACC 6h | 938.448 | 108.253 | 0.783508 | 0.162794 |
| BABA 6h | 1133.25 | 95.5252 | 0.97447 | 0.0563588 |
| Chitin 6h | 945.959 | 54.9225 | 0.859015 | 0.0497168 |
| Epi 6h | 1375.05 | 194.514 | 1.16054 | 0.154213 |
| SA 6h | 1274.22 | 341.066 | 1.16643 | 0.216762 |
| Me-JA 6h | 974.927 | 380.7 | 0.82632 | 0.254523 |
Source Transcript PGSC0003DMT400042159 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G47890.1 | +1 | 4e-62 | 218 | 157/425 (37%) | receptor like protein 7 | chr1:17643976-17647035 FORWARD LENGTH=1019 |
