Probe CUST_18247_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18247_PI426222305 | JHI_St_60k_v1 | DMT400008936 | CCAAATTCCTATATTGTTTTATCAGCAGATGCTTTCAGGGCATGGACAAATGATAGTTTG |
All Microarray Probes Designed to Gene DMG402003479
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18378_PI426222305 | JHI_St_60k_v1 | DMT400008935 | GTTGCCCTTTCTCTGTGAAAAGAAATCACTAATACTTTTTATGTGTCGATGTTGTGGATT |
| CUST_18247_PI426222305 | JHI_St_60k_v1 | DMT400008936 | CCAAATTCCTATATTGTTTTATCAGCAGATGCTTTCAGGGCATGGACAAATGATAGTTTG |
Microarray Signals from CUST_18247_PI426222305
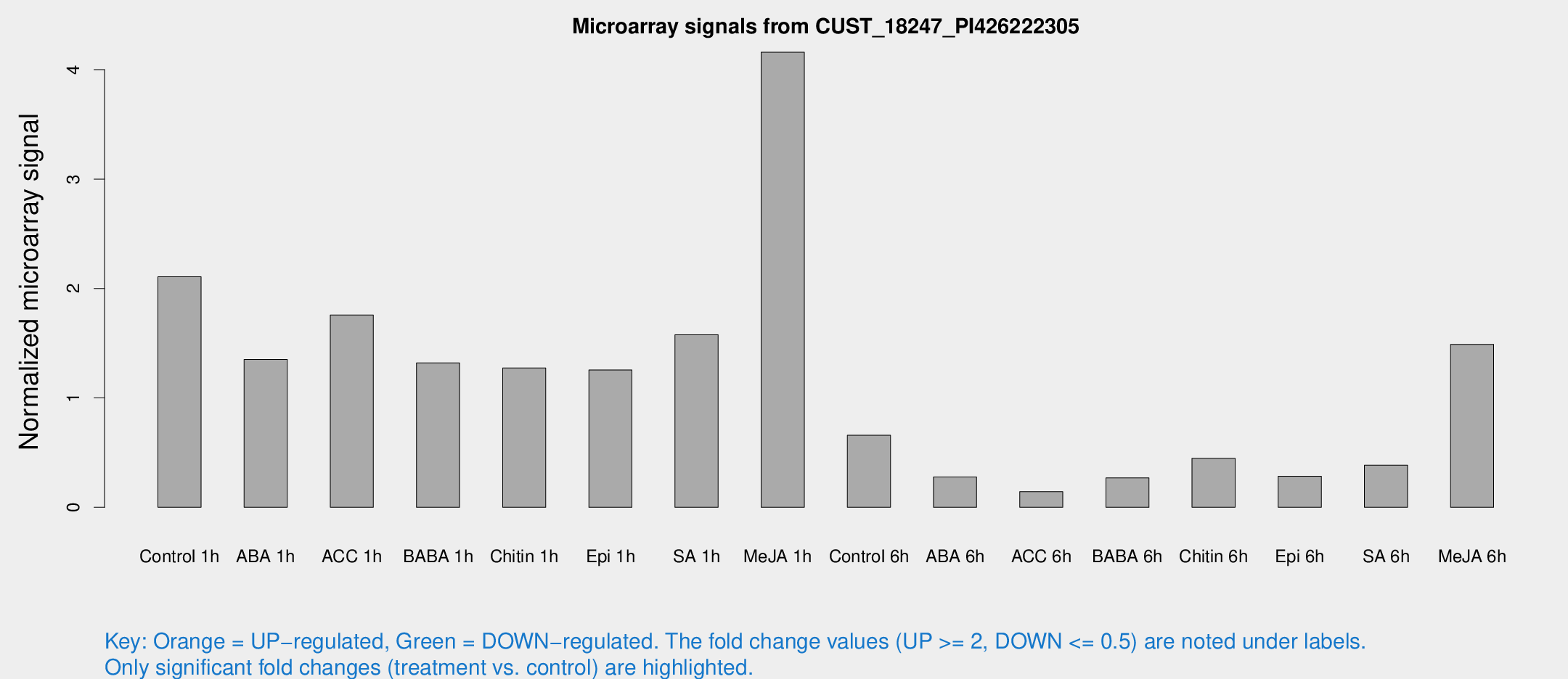
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4507.75 | 260.957 | 2.10794 | 0.249809 |
| ABA 1h | 2792.11 | 782.606 | 1.35209 | 0.418358 |
| ACC 1h | 4325.01 | 1266.82 | 1.75689 | 0.544691 |
| BABA 1h | 2724.51 | 179.915 | 1.31943 | 0.184722 |
| Chitin 1h | 2560.29 | 551.226 | 1.27351 | 0.252308 |
| Epi 1h | 2329.42 | 180.979 | 1.25463 | 0.104985 |
| SA 1h | 4540.85 | 2384.79 | 1.57762 | 1.05562 |
| Me-JA 1h | 7313.86 | 727.932 | 4.15963 | 0.417525 |
| Control 6h | 1627.91 | 635.61 | 0.659043 | 0.21097 |
| ABA 6h | 636.494 | 68.4096 | 0.277658 | 0.0440125 |
| ACC 6h | 365.307 | 68.8407 | 0.143301 | 0.0343148 |
| BABA 6h | 840.514 | 333.597 | 0.268331 | 0.231314 |
| Chitin 6h | 1020.41 | 72.8621 | 0.447277 | 0.0359105 |
| Epi 6h | 949.165 | 430.129 | 0.284502 | 0.313237 |
| SA 6h | 881.687 | 221.72 | 0.385526 | 0.0724287 |
| Me-JA 6h | 3416.66 | 839.021 | 1.48932 | 0.490365 |
Source Transcript PGSC0003DMT400008936 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52820.1 | +3 | 3e-62 | 205 | 118/310 (38%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr1:19669216-19670321 FORWARD LENGTH=317 |
