Probe CUST_18229_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18229_PI426222305 | JHI_St_60k_v1 | DMT400055037 | ATGTTGTTGCAACAATGGTGTCACTTACACATCTTTGGCTTCATGGGAACCAATTTTCAG |
All Microarray Probes Designed to Gene DMG400021353
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18352_PI426222305 | JHI_St_60k_v1 | DMT400055038 | AAACTTGCTCCTGATAAAGAGAAATCTGTGGTCACCAGGCTGGCCGGAACTTTTGGATAT |
| CUST_18229_PI426222305 | JHI_St_60k_v1 | DMT400055037 | ATGTTGTTGCAACAATGGTGTCACTTACACATCTTTGGCTTCATGGGAACCAATTTTCAG |
Microarray Signals from CUST_18229_PI426222305
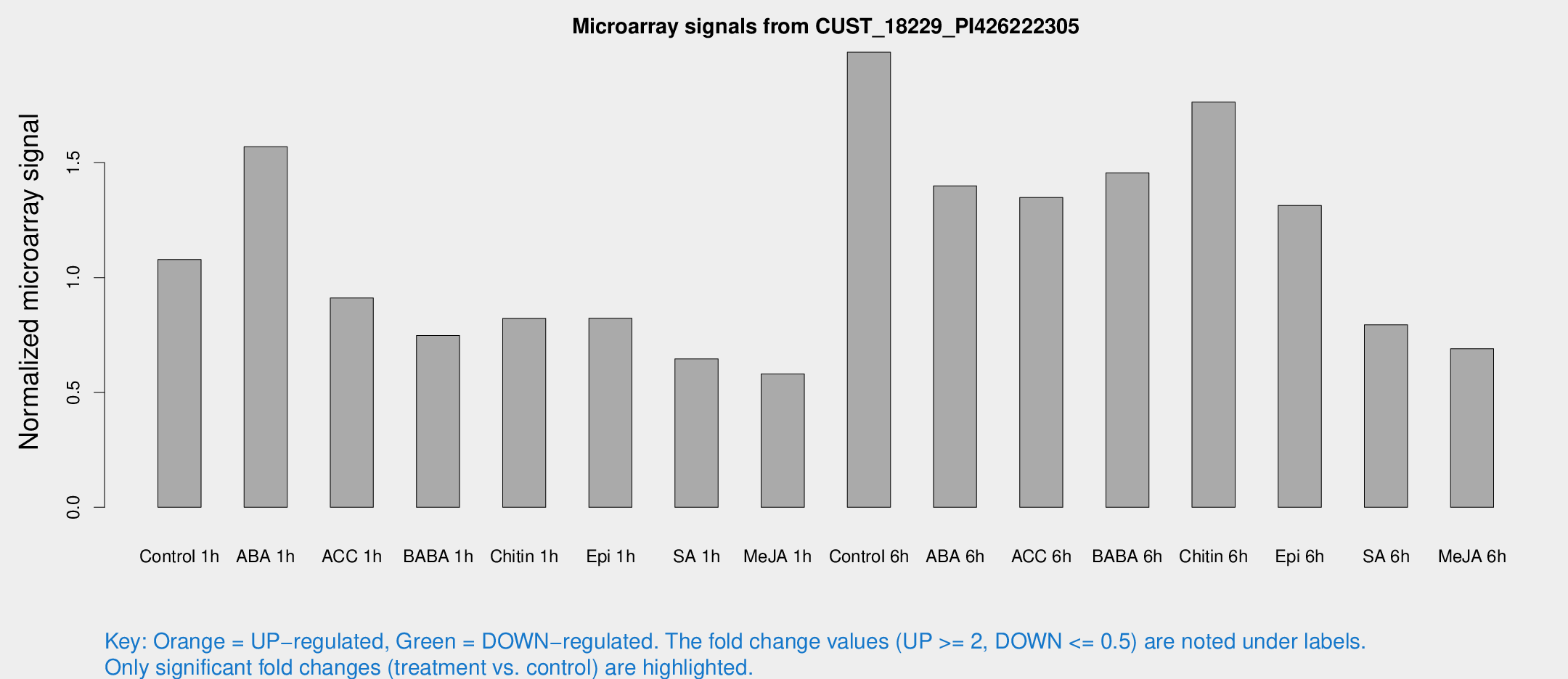
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 62.3051 | 11.4997 | 1.0787 | 0.180682 |
| ABA 1h | 84.7739 | 25.2617 | 1.56982 | 0.470285 |
| ACC 1h | 54.3473 | 10.2523 | 0.911064 | 0.122092 |
| BABA 1h | 40.2184 | 4.4268 | 0.747914 | 0.0823524 |
| Chitin 1h | 46.6743 | 15.0657 | 0.822227 | 0.290189 |
| Epi 1h | 39.9722 | 4.11829 | 0.823069 | 0.0841104 |
| SA 1h | 37.564 | 4.57202 | 0.645912 | 0.0737503 |
| Me-JA 1h | 26.632 | 4.08185 | 0.580599 | 0.089265 |
| Control 6h | 126.13 | 40.6375 | 1.98092 | 0.63506 |
| ABA 6h | 85.1414 | 13.3293 | 1.39911 | 0.165732 |
| ACC 6h | 97.5256 | 29.0128 | 1.34858 | 0.430567 |
| BABA 6h | 103.295 | 32.1546 | 1.45583 | 0.549054 |
| Chitin 6h | 114.955 | 31.7364 | 1.76354 | 0.637378 |
| Epi 6h | 100.397 | 34.8569 | 1.31394 | 0.943506 |
| SA 6h | 80.8032 | 42.6682 | 0.794321 | 2.04358 |
| Me-JA 6h | 56.0943 | 30.9002 | 0.690302 | 0.513635 |
Source Transcript PGSC0003DMT400055037 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01820.1 | +2 | 0.0 | 569 | 317/621 (51%) | Leucine-rich repeat protein kinase family protein | chr2:357664-360681 REVERSE LENGTH=943 |
