Probe CUST_17781_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17781_PI426222305 | JHI_St_60k_v1 | DMT400066826 | CCTTGCTGCATATATTCTTCCATTGCTCCATGCCTACTTCCAAAAACAAAATCACTTTTT |
All Microarray Probes Designed to Gene DMG400025981
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17781_PI426222305 | JHI_St_60k_v1 | DMT400066826 | CCTTGCTGCATATATTCTTCCATTGCTCCATGCCTACTTCCAAAAACAAAATCACTTTTT |
| CUST_17758_PI426222305 | JHI_St_60k_v1 | DMT400066819 | CTGCTAGCACTTGCTTCTGGAGCACCCAATACTCGGTCGTTGCAGAATATTGTAATTCAA |
Microarray Signals from CUST_17781_PI426222305
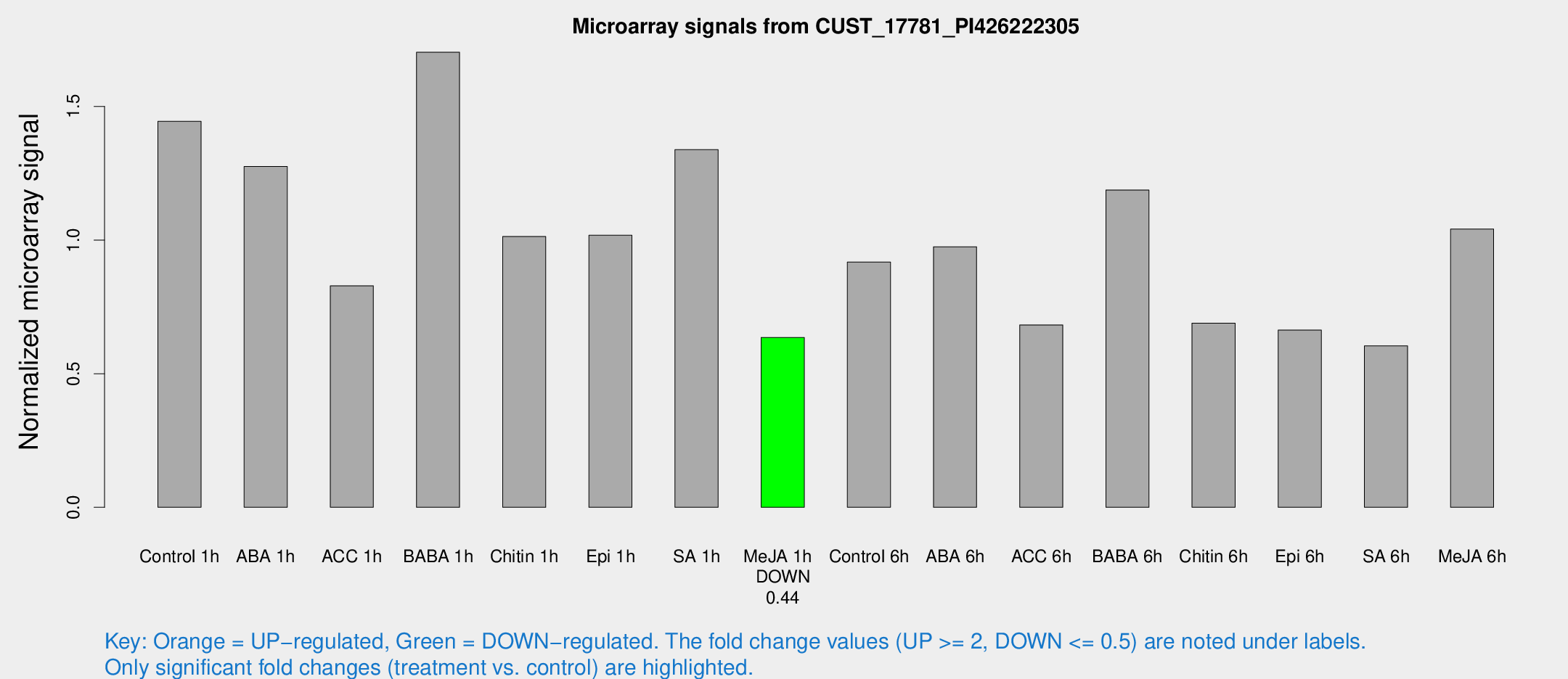
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 52.4858 | 8.57167 | 1.44465 | 0.206061 |
| ABA 1h | 41.5589 | 8.6972 | 1.27524 | 0.261176 |
| ACC 1h | 31.9859 | 7.04453 | 0.828661 | 0.153098 |
| BABA 1h | 58.6504 | 5.68869 | 1.70306 | 0.138596 |
| Chitin 1h | 34.064 | 8.0248 | 1.01329 | 0.301427 |
| Epi 1h | 31.2306 | 3.5779 | 1.01794 | 0.117775 |
| SA 1h | 51.3377 | 11.6833 | 1.33828 | 0.387246 |
| Me-JA 1h | 19.4927 | 4.6296 | 0.635779 | 0.124004 |
| Control 6h | 38.297 | 13.1619 | 0.9179 | 0.340791 |
| ABA 6h | 38.3909 | 7.84121 | 0.975088 | 0.165151 |
| ACC 6h | 28.3423 | 4.31642 | 0.682422 | 0.213225 |
| BABA 6h | 50.4196 | 11.814 | 1.18735 | 0.301259 |
| Chitin 6h | 30.3765 | 9.93876 | 0.689056 | 0.350856 |
| Epi 6h | 36.6217 | 17.6298 | 0.663192 | 0.644595 |
| SA 6h | 31.0692 | 17.9521 | 0.603981 | 0.735699 |
| Me-JA 6h | 44.7327 | 20.3076 | 1.04135 | 0.411786 |
Source Transcript PGSC0003DMT400066826 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G44980.2 | +1 | 3e-130 | 427 | 249/412 (60%) | SNF2 domain-containing protein / helicase domain-containing protein | chr2:18552343-18556669 REVERSE LENGTH=877 |
