Probe CUST_17758_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17758_PI426222305 | JHI_St_60k_v1 | DMT400066819 | CTGCTAGCACTTGCTTCTGGAGCACCCAATACTCGGTCGTTGCAGAATATTGTAATTCAA |
All Microarray Probes Designed to Gene DMG400025981
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17781_PI426222305 | JHI_St_60k_v1 | DMT400066826 | CCTTGCTGCATATATTCTTCCATTGCTCCATGCCTACTTCCAAAAACAAAATCACTTTTT |
| CUST_17758_PI426222305 | JHI_St_60k_v1 | DMT400066819 | CTGCTAGCACTTGCTTCTGGAGCACCCAATACTCGGTCGTTGCAGAATATTGTAATTCAA |
Microarray Signals from CUST_17758_PI426222305
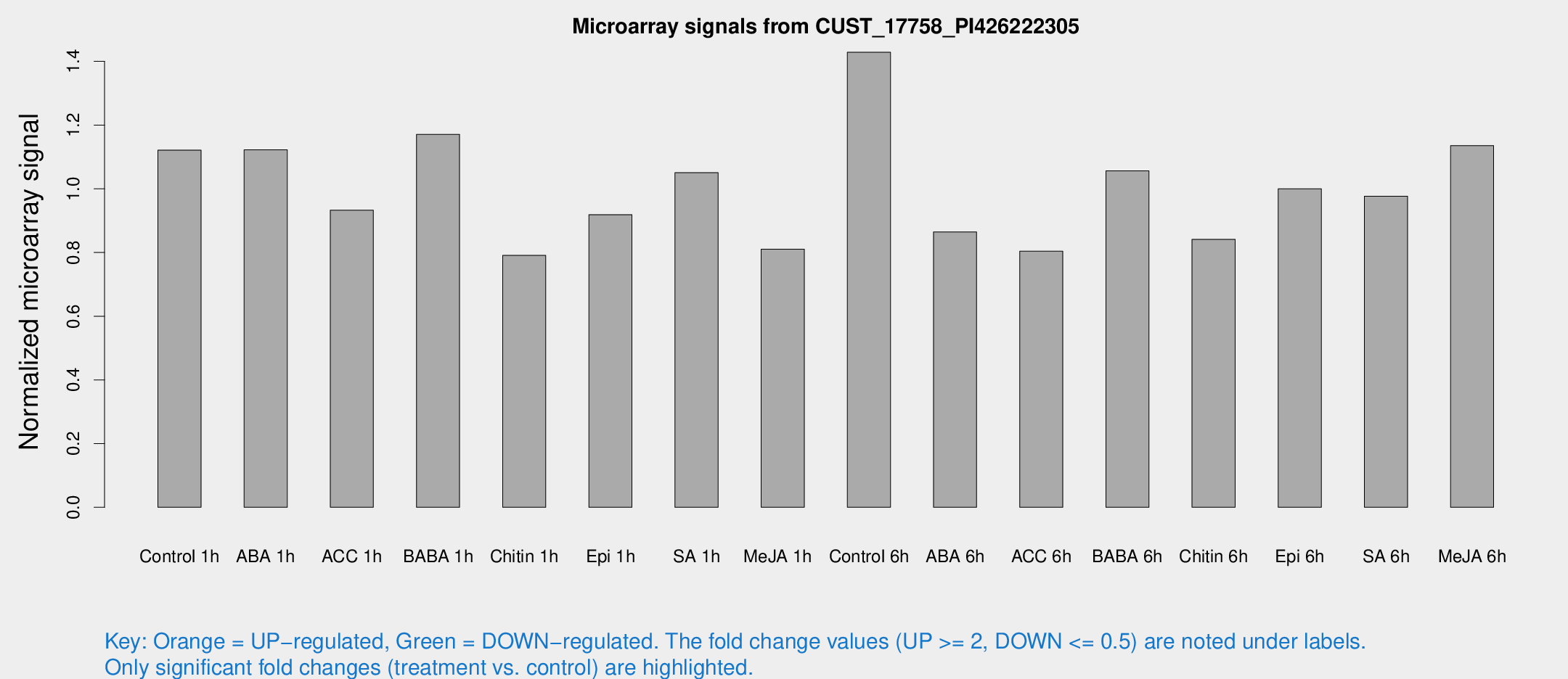
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 73.5347 | 18.279 | 1.12142 | 0.251174 |
| ABA 1h | 62.3132 | 10.6981 | 1.1226 | 0.239582 |
| ACC 1h | 64.5642 | 17.4648 | 0.933068 | 0.255359 |
| BABA 1h | 71.6512 | 14.1831 | 1.17096 | 0.140903 |
| Chitin 1h | 44.3636 | 6.34903 | 0.790784 | 0.0716443 |
| Epi 1h | 49.9127 | 8.4114 | 0.918865 | 0.183963 |
| SA 1h | 68.0956 | 12.5053 | 1.0506 | 0.173942 |
| Me-JA 1h | 41.7436 | 7.8203 | 0.810592 | 0.0792155 |
| Control 6h | 95.4722 | 24.6804 | 1.42874 | 0.331249 |
| ABA 6h | 59.1857 | 12.6057 | 0.864738 | 0.171123 |
| ACC 6h | 57.538 | 8.33327 | 0.803864 | 0.13199 |
| BABA 6h | 81.1042 | 23.463 | 1.05636 | 0.383068 |
| Chitin 6h | 55.8334 | 7.81518 | 0.841412 | 0.135378 |
| Epi 6h | 79.9223 | 31.7352 | 1.00014 | 0.454498 |
| SA 6h | 64.0492 | 16.3731 | 0.976861 | 0.435532 |
| Me-JA 6h | 75.5413 | 22.3139 | 1.13557 | 0.259658 |
Source Transcript PGSC0003DMT400066819 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G44980.2 | +3 | 3e-162 | 483 | 246/361 (68%) | SNF2 domain-containing protein / helicase domain-containing protein | chr2:18552343-18556669 REVERSE LENGTH=877 |
