Probe CUST_17585_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17585_PI426222305 | JHI_St_60k_v1 | DMT400001588 | AGACAAATTATACTTTTAACCCGTTACTAGTTGGAGAGTGACACTTTGGAGCAACAGCTT |
All Microarray Probes Designed to Gene DMG400000587
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17585_PI426222305 | JHI_St_60k_v1 | DMT400001588 | AGACAAATTATACTTTTAACCCGTTACTAGTTGGAGAGTGACACTTTGGAGCAACAGCTT |
| CUST_17468_PI426222305 | JHI_St_60k_v1 | DMT400001587 | GCTACAGAGCTGCCAATCCCTTTGATATTGACACTCTTTAAAATTGTAAATCAGTATCAT |
Microarray Signals from CUST_17585_PI426222305
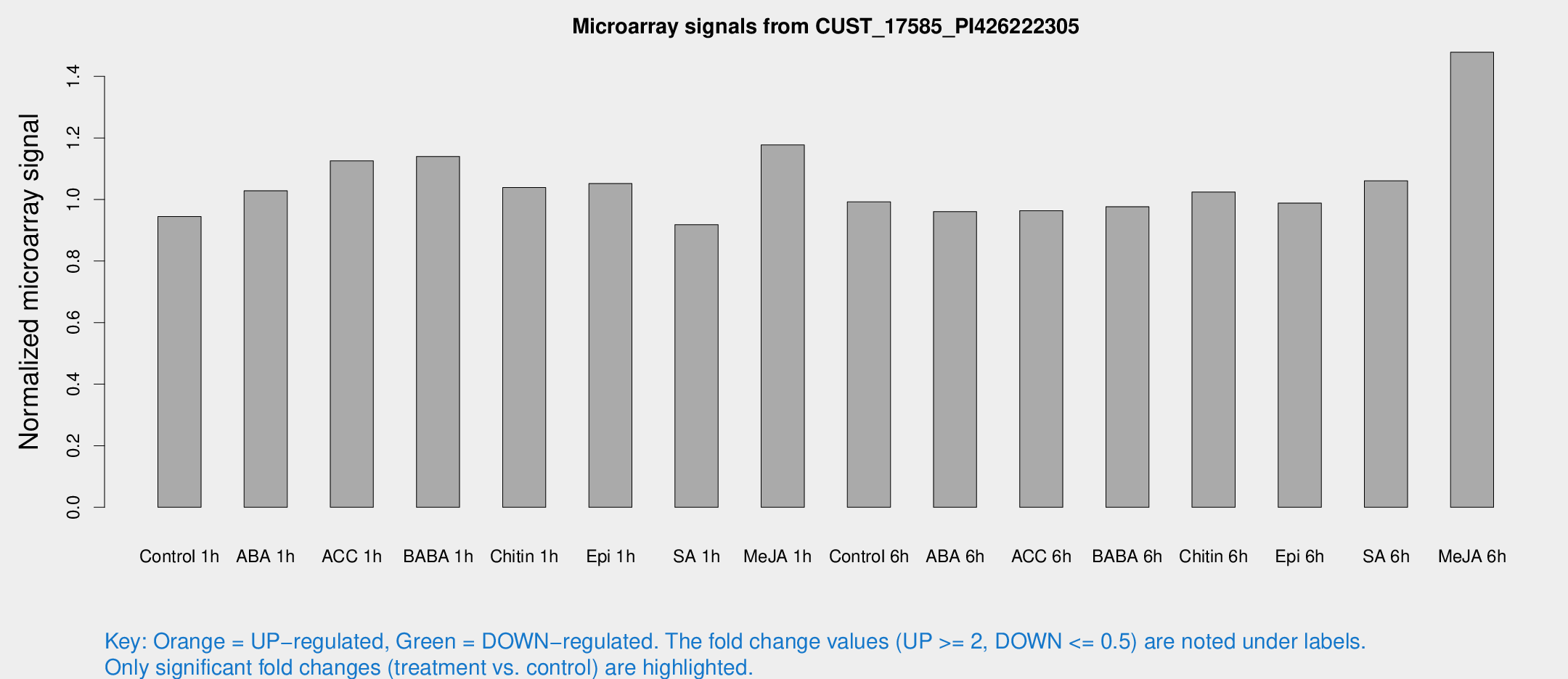
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.03452 | 3.50867 | 0.94486 | 0.547458 |
| ABA 1h | 5.79286 | 3.35641 | 1.02836 | 0.595517 |
| ACC 1h | 7.54828 | 4.48997 | 1.12581 | 0.652135 |
| BABA 1h | 7.13829 | 3.78946 | 1.13971 | 0.615956 |
| Chitin 1h | 5.99286 | 3.49836 | 1.03888 | 0.601683 |
| Epi 1h | 5.81758 | 3.38279 | 1.05167 | 0.609077 |
| SA 1h | 6.01837 | 3.49337 | 0.918406 | 0.531842 |
| Me-JA 1h | 6.16869 | 3.6093 | 1.17708 | 0.68228 |
| Control 6h | 6.34253 | 3.68071 | 0.992114 | 0.574529 |
| ABA 6h | 6.50749 | 3.7719 | 0.960792 | 0.556356 |
| ACC 6h | 7.21071 | 4.28352 | 0.963724 | 0.558064 |
| BABA 6h | 6.97621 | 4.05075 | 0.976591 | 0.565525 |
| Chitin 6h | 6.95198 | 4.032 | 1.02442 | 0.593161 |
| Epi 6h | 7.21076 | 4.25255 | 0.988415 | 0.572789 |
| SA 6h | 6.68018 | 3.87066 | 1.06082 | 0.614517 |
| Me-JA 6h | 10.9828 | 4.59577 | 1.47843 | 0.683605 |
Source Transcript PGSC0003DMT400001588 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G54180.1 | +1 | 3e-158 | 472 | 265/482 (55%) | plastid transcriptionally active 15 | chr5:21988544-21990183 FORWARD LENGTH=500 |
