Probe CUST_17468_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17468_PI426222305 | JHI_St_60k_v1 | DMT400001587 | GCTACAGAGCTGCCAATCCCTTTGATATTGACACTCTTTAAAATTGTAAATCAGTATCAT |
All Microarray Probes Designed to Gene DMG400000587
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17585_PI426222305 | JHI_St_60k_v1 | DMT400001588 | AGACAAATTATACTTTTAACCCGTTACTAGTTGGAGAGTGACACTTTGGAGCAACAGCTT |
| CUST_17468_PI426222305 | JHI_St_60k_v1 | DMT400001587 | GCTACAGAGCTGCCAATCCCTTTGATATTGACACTCTTTAAAATTGTAAATCAGTATCAT |
Microarray Signals from CUST_17468_PI426222305
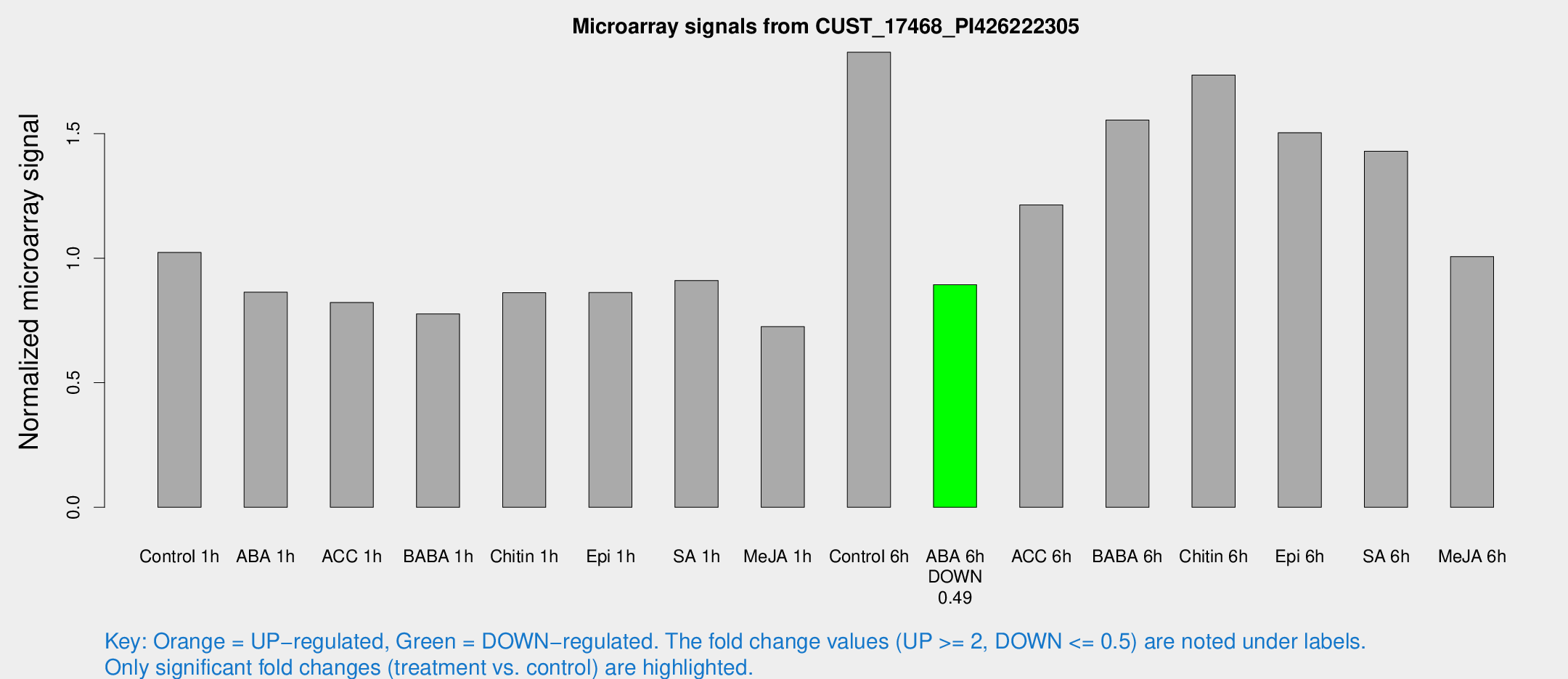
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1624.46 | 265.542 | 1.0228 | 0.122263 |
| ABA 1h | 1190.86 | 127.05 | 0.86321 | 0.0498894 |
| ACC 1h | 1362.04 | 267.395 | 0.821728 | 0.120437 |
| BABA 1h | 1208.14 | 236.46 | 0.776489 | 0.0927582 |
| Chitin 1h | 1205.49 | 117.434 | 0.861338 | 0.0663137 |
| Epi 1h | 1163.93 | 132.729 | 0.861979 | 0.072044 |
| SA 1h | 1456.08 | 152.078 | 0.910278 | 0.0525936 |
| Me-JA 1h | 926.033 | 115.754 | 0.725287 | 0.0419539 |
| Control 6h | 2960.94 | 627.284 | 1.82654 | 0.280526 |
| ABA 6h | 1469.71 | 91.7684 | 0.893627 | 0.0516358 |
| ACC 6h | 2226.65 | 416.89 | 1.21364 | 0.117196 |
| BABA 6h | 2784.41 | 566.077 | 1.55412 | 0.291959 |
| Chitin 6h | 2937.37 | 543.231 | 1.73476 | 0.25464 |
| Epi 6h | 2622.41 | 162.016 | 1.50366 | 0.138422 |
| SA 6h | 2219.18 | 288.572 | 1.4289 | 0.0825322 |
| Me-JA 6h | 1645.22 | 393.487 | 1.00652 | 0.176724 |
Source Transcript PGSC0003DMT400001587 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G54180.1 | +2 | 1e-155 | 466 | 247/413 (60%) | plastid transcriptionally active 15 | chr5:21988544-21990183 FORWARD LENGTH=500 |
