Probe CUST_17377_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17377_PI426222305 | JHI_St_60k_v1 | DMT400013832 | GGCTCGAATTTTAATTAAGGAAGGAACAAATACTGCACCACATTTCTAAGCAGAACAATT |
All Microarray Probes Designed to Gene DMG400005410
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17364_PI426222305 | JHI_St_60k_v1 | DMT400013831 | TAAAAATAACCAAAGTGGCGATGGCGACTGAAGCAGTAATCAAGGTAGTAGCACTCTGCG |
| CUST_17377_PI426222305 | JHI_St_60k_v1 | DMT400013832 | GGCTCGAATTTTAATTAAGGAAGGAACAAATACTGCACCACATTTCTAAGCAGAACAATT |
Microarray Signals from CUST_17377_PI426222305
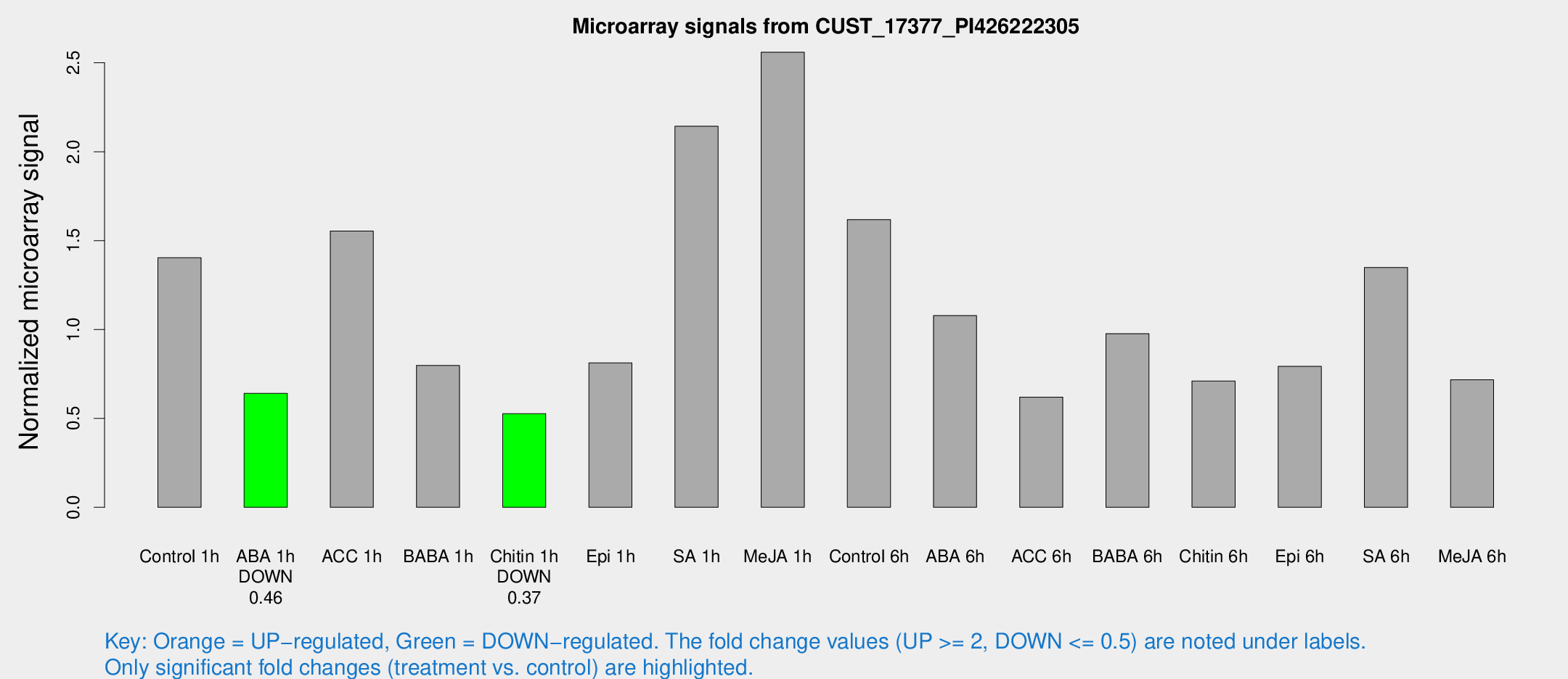
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.2691 | 3.51634 | 1.40425 | 0.290933 |
| ABA 1h | 7.25155 | 3.33225 | 0.640836 | 0.30251 |
| ACC 1h | 24.043 | 10.4731 | 1.55342 | 0.63852 |
| BABA 1h | 10.8205 | 3.69354 | 0.797588 | 0.341805 |
| Chitin 1h | 5.98953 | 3.48158 | 0.526194 | 0.304712 |
| Epi 1h | 9.2521 | 3.43034 | 0.812677 | 0.331505 |
| SA 1h | 36.1037 | 14.1366 | 2.14328 | 1.39294 |
| Me-JA 1h | 27.5781 | 6.15105 | 2.55962 | 0.38494 |
| Control 6h | 21.5134 | 4.48018 | 1.61852 | 0.343441 |
| ABA 6h | 16.0773 | 5.19984 | 1.07819 | 0.326038 |
| ACC 6h | 9.21026 | 4.42013 | 0.61968 | 0.305568 |
| BABA 6h | 16.2621 | 6.23356 | 0.976512 | 0.431117 |
| Chitin 6h | 10.7233 | 4.04666 | 0.710038 | 0.33567 |
| Epi 6h | 14.0519 | 6.78032 | 0.792679 | 0.404035 |
| SA 6h | 19.9327 | 6.60167 | 1.34867 | 0.491012 |
| Me-JA 6h | 9.9632 | 3.64145 | 0.71721 | 0.321303 |
Source Transcript PGSC0003DMT400013832 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G27890.1 | +2 | 4e-77 | 238 | 112/165 (68%) | NADPH:quinone oxidoreductase | chr3:10350807-10351938 REVERSE LENGTH=196 |
