Probe CUST_16715_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16715_PI426222305 | JHI_St_60k_v1 | DMT400069229 | GGAAATGCTTGCACTTTTGTTCTATGACTATGAGGTACACGAATAAAACAAGAAACTGTT |
All Microarray Probes Designed to Gene DMG400026930
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16715_PI426222305 | JHI_St_60k_v1 | DMT400069229 | GGAAATGCTTGCACTTTTGTTCTATGACTATGAGGTACACGAATAAAACAAGAAACTGTT |
Microarray Signals from CUST_16715_PI426222305
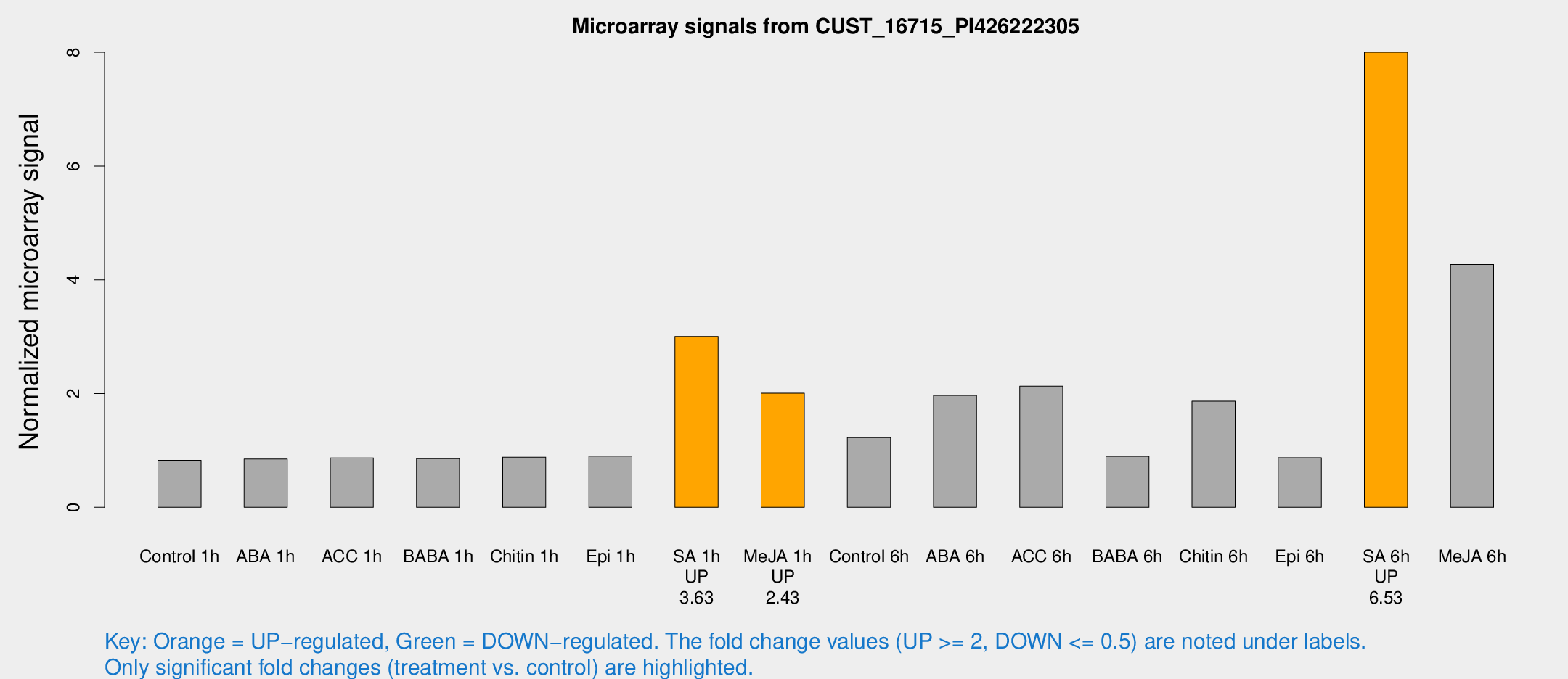
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.5777 | 3.06843 | 0.826981 | 0.453837 |
| ABA 1h | 5.02544 | 2.91136 | 0.848711 | 0.486597 |
| ACC 1h | 6.08588 | 3.56588 | 0.86837 | 0.503456 |
| BABA 1h | 5.58237 | 3.24169 | 0.855245 | 0.495633 |
| Chitin 1h | 5.2149 | 3.04586 | 0.882168 | 0.495711 |
| Epi 1h | 5.15337 | 2.99805 | 0.899026 | 0.510531 |
| SA 1h | 22.8487 | 7.24451 | 3.0056 | 0.761328 |
| Me-JA 1h | 11.105 | 3.16681 | 2.00633 | 0.573591 |
| Control 6h | 8.62522 | 3.15982 | 1.22514 | 0.496491 |
| ABA 6h | 17.2372 | 6.08438 | 1.96777 | 1.02573 |
| ACC 6h | 18.3518 | 5.08401 | 2.13119 | 1.24511 |
| BABA 6h | 6.98906 | 3.49049 | 0.896858 | 0.467326 |
| Chitin 6h | 13.4543 | 3.53274 | 1.86744 | 0.490862 |
| Epi 6h | 6.88904 | 3.63383 | 0.872697 | 0.469768 |
| SA 6h | 55.296 | 10.506 | 8.00133 | 1.28107 |
| Me-JA 6h | 46.0476 | 20.331 | 4.27134 | 6.86574 |
Source Transcript PGSC0003DMT400069229 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36780.1 | +1 | 2e-173 | 506 | 247/482 (51%) | UDP-Glycosyltransferase superfamily protein | chr2:15417618-15419108 REVERSE LENGTH=496 |
