Probe CUST_16614_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16614_PI426222305 | JHI_St_60k_v1 | DMT400069218 | GCTACTGATCGATCGAGCAGGCTATCTAATAAATAATGTGTTAGGAGATTTGTACAATAA |
All Microarray Probes Designed to Gene DMG401026923
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16614_PI426222305 | JHI_St_60k_v1 | DMT400069218 | GCTACTGATCGATCGAGCAGGCTATCTAATAAATAATGTGTTAGGAGATTTGTACAATAA |
| CUST_16521_PI426222305 | JHI_St_60k_v1 | DMT400069219 | GGTGATCAATTGGAGGTAAAACATGACTCTTCTATTTCATAACTATTCCATCTATCGCAA |
Microarray Signals from CUST_16614_PI426222305
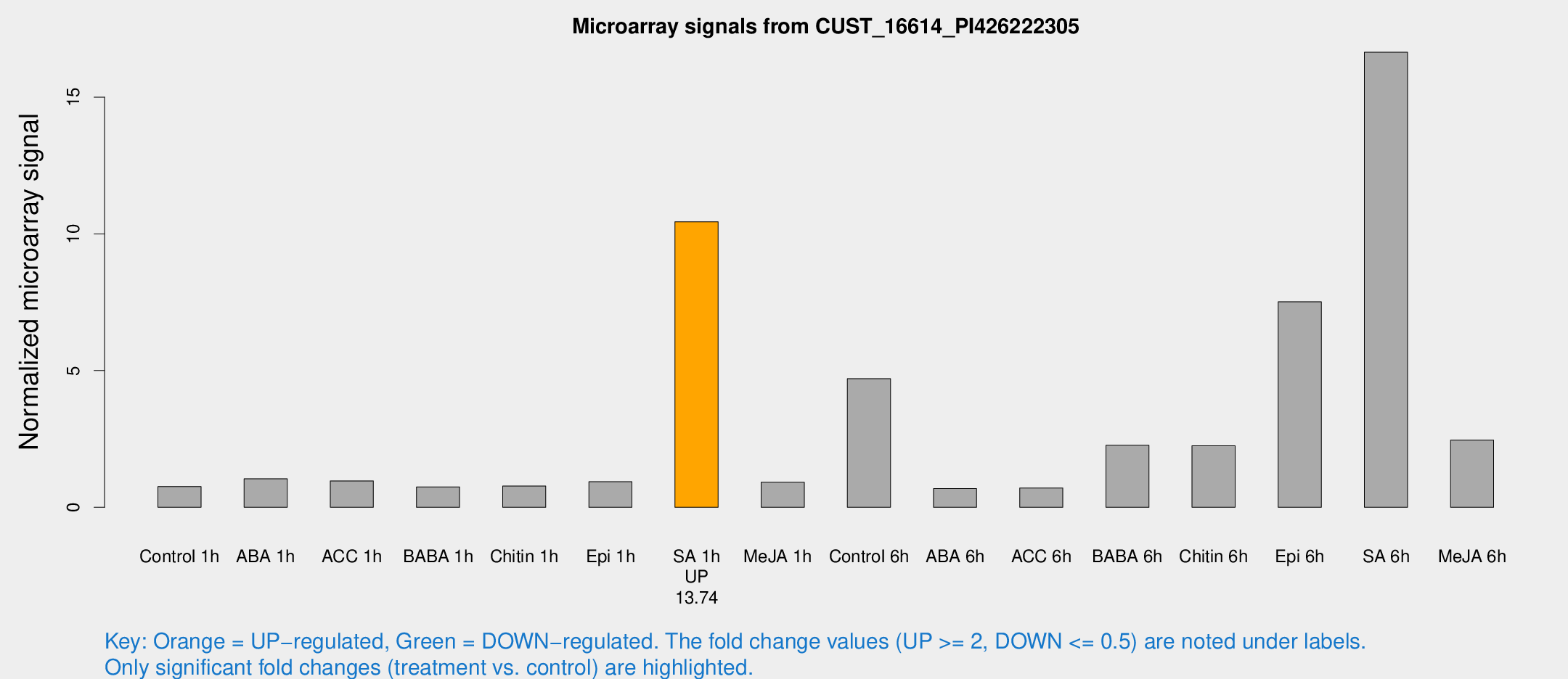
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.5757 | 2.86011 | 0.759613 | 0.391504 |
| ABA 1h | 7.00654 | 2.78475 | 1.04424 | 0.449431 |
| ACC 1h | 7.42958 | 3.13296 | 0.96678 | 0.425876 |
| BABA 1h | 5.30028 | 3.03069 | 0.745079 | 0.425734 |
| Chitin 1h | 4.95395 | 2.88144 | 0.778147 | 0.431764 |
| Epi 1h | 5.96809 | 2.88213 | 0.935092 | 0.460628 |
| SA 1h | 79.8441 | 7.54939 | 10.4405 | 1.16855 |
| Me-JA 1h | 5.52393 | 2.94805 | 0.916544 | 0.488556 |
| Control 6h | 47.6582 | 23.8364 | 4.70495 | 2.91415 |
| ABA 6h | 5.35504 | 3.10594 | 0.682529 | 0.395313 |
| ACC 6h | 6.10824 | 3.61966 | 0.70681 | 0.409605 |
| BABA 6h | 22.1054 | 8.40833 | 2.26939 | 1.23591 |
| Chitin 6h | 21.1739 | 8.25454 | 2.25172 | 1.11145 |
| Epi 6h | 71.8078 | 22.4554 | 7.51803 | 4.24053 |
| SA 6h | 127.705 | 28.2102 | 16.6426 | 2.30599 |
| Me-JA 6h | 30.0603 | 21.0034 | 2.45864 | 2.14981 |
Source Transcript PGSC0003DMT400069218 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G24530.1 | +2 | 3e-176 | 502 | 231/338 (68%) | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein | chr5:8378964-8383154 FORWARD LENGTH=341 |
