Probe CUST_16411_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16411_PI426222305 | JHI_St_60k_v1 | DMT400089399 | CGGATTGTCTTCATGAGGAGGTAATTGAAGCTGCAGATATAGGTTACGCGATTGCAGTTG |
All Microarray Probes Designed to Gene DMG400038970
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16411_PI426222305 | JHI_St_60k_v1 | DMT400089399 | CGGATTGTCTTCATGAGGAGGTAATTGAAGCTGCAGATATAGGTTACGCGATTGCAGTTG |
Microarray Signals from CUST_16411_PI426222305
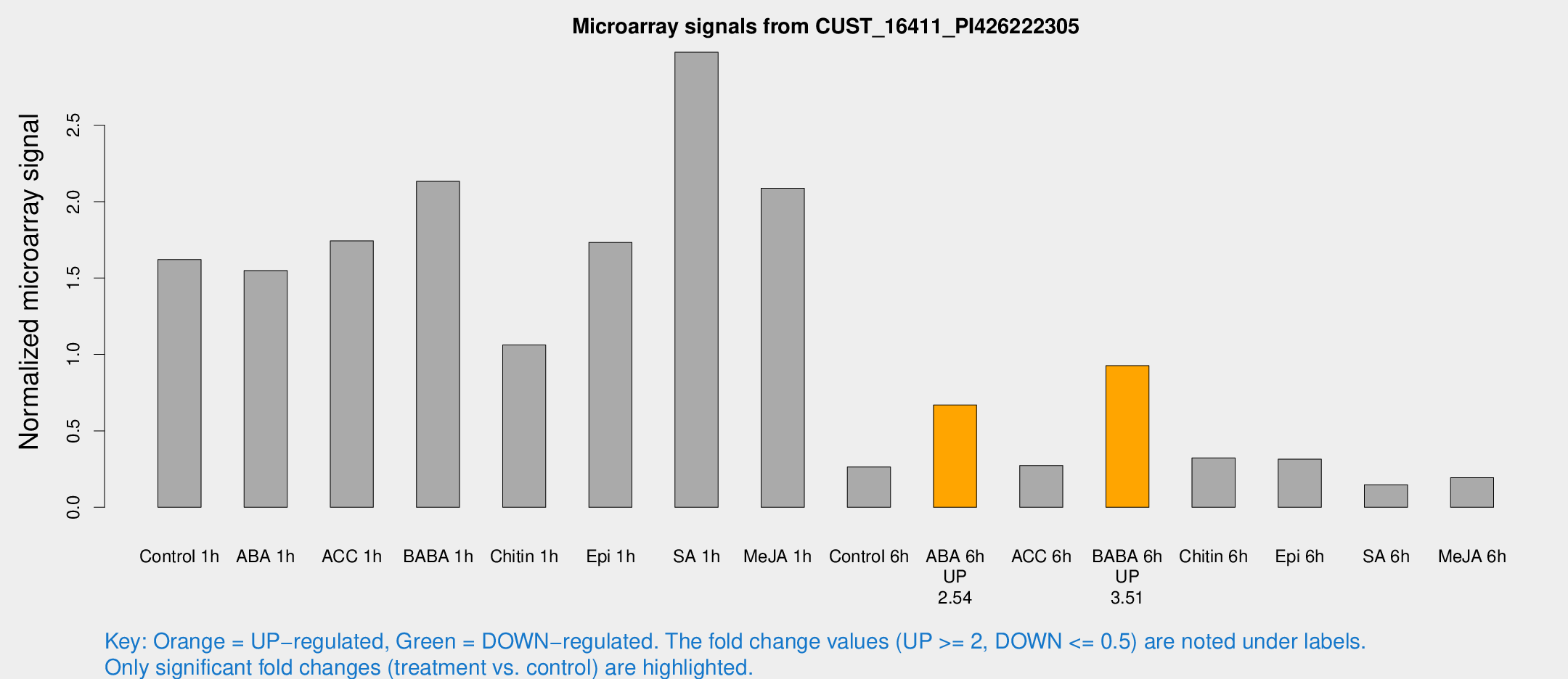
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 213.7 | 38.6982 | 1.62107 | 0.180877 |
| ABA 1h | 179.16 | 28.6686 | 1.5482 | 0.135821 |
| ACC 1h | 248.952 | 63.1824 | 1.74272 | 0.421536 |
| BABA 1h | 269.073 | 41.9819 | 2.13241 | 0.176657 |
| Chitin 1h | 127.994 | 29.1759 | 1.06195 | 0.178988 |
| Epi 1h | 196.351 | 28.8459 | 1.7331 | 0.333513 |
| SA 1h | 410.617 | 83.2749 | 2.97702 | 0.544717 |
| Me-JA 1h | 217.797 | 13.0078 | 2.08676 | 0.138857 |
| Control 6h | 36.6057 | 9.08976 | 0.264079 | 0.0552017 |
| ABA 6h | 92.2923 | 11.5922 | 0.669682 | 0.087893 |
| ACC 6h | 41.2805 | 7.0019 | 0.273622 | 0.031574 |
| BABA 6h | 133.714 | 12.9596 | 0.926829 | 0.0741087 |
| Chitin 6h | 44.6817 | 6.04957 | 0.322889 | 0.0365017 |
| Epi 6h | 46.1617 | 6.41149 | 0.314815 | 0.0439732 |
| SA 6h | 19.1378 | 3.79199 | 0.147847 | 0.0310925 |
| Me-JA 6h | 25.418 | 4.25741 | 0.193895 | 0.0305718 |
Source Transcript PGSC0003DMT400089399 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22560.1 | +1 | 8e-41 | 138 | 76/160 (48%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr3:7998915-7999442 REVERSE LENGTH=175 |
