Probe CUST_16264_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16264_PI426222305 | JHI_St_60k_v1 | DMT400016212 | GTACAGTGACATTACAAGGCGAGTTCAAGAAATATGGAGATAAGGTTGATTTAGTAGTTT |
All Microarray Probes Designed to Gene DMG400006337
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16264_PI426222305 | JHI_St_60k_v1 | DMT400016212 | GTACAGTGACATTACAAGGCGAGTTCAAGAAATATGGAGATAAGGTTGATTTAGTAGTTT |
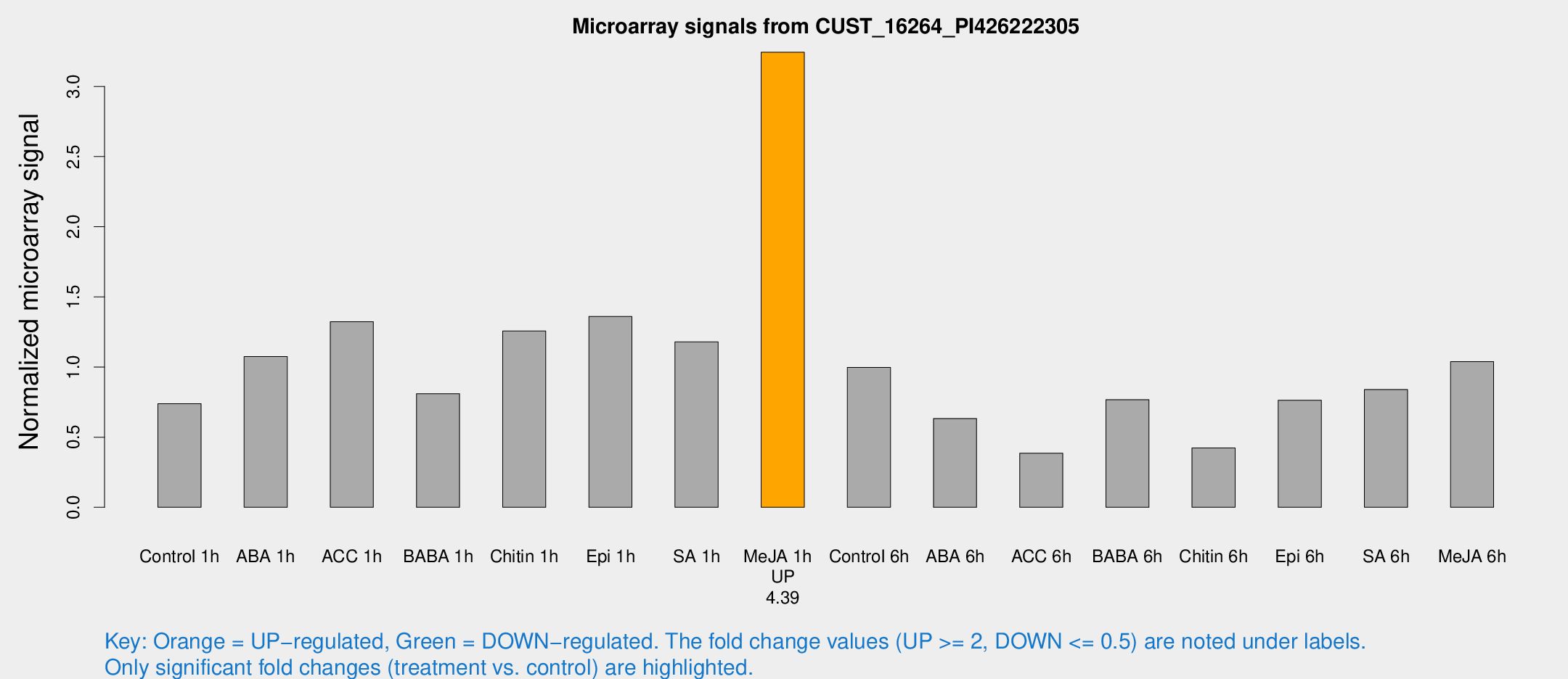
Microarray Signals from CUST_16264_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14.9561 | 4.44844 | 0.738495 | 0.266125 |
| ABA 1h | 18.6981 | 5.11616 | 1.07461 | 0.308316 |
| ACC 1h | 24.4448 | 4.16528 | 1.32337 | 0.228068 |
| BABA 1h | 15.7207 | 4.75749 | 0.810157 | 0.427136 |
| Chitin 1h | 20.2137 | 3.99274 | 1.25683 | 0.247909 |
| Epi 1h | 22.0983 | 5.23894 | 1.36053 | 0.33872 |
| SA 1h | 22.5667 | 4.55699 | 1.17932 | 0.226006 |
| Me-JA 1h | 47.3469 | 4.47887 | 3.24399 | 0.306104 |
| Control 6h | 19.2505 | 5.1484 | 0.99769 | 0.250299 |
| ABA 6h | 14.08 | 5.86054 | 0.632927 | 0.314677 |
| ACC 6h | 8.05407 | 4.75329 | 0.386008 | 0.222642 |
| BABA 6h | 15.5922 | 4.23175 | 0.767395 | 0.211433 |
| Chitin 6h | 8.19467 | 4.23349 | 0.422637 | 0.224459 |
| Epi 6h | 16.7763 | 5.05957 | 0.763575 | 0.277682 |
| SA 6h | 17.6267 | 7.13561 | 0.839773 | 0.559662 |
| Me-JA 6h | 24.6814 | 11.2043 | 1.0383 | 0.643595 |
Source Transcript PGSC0003DMT400016212 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22560.1 | +3 | 6e-39 | 135 | 73/159 (46%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr3:7998915-7999442 REVERSE LENGTH=175 |
