Probe CUST_16082_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16082_PI426222305 | JHI_St_60k_v1 | DMT400040040 | GACCTGATAATGAAATCAGCTTGATCCAAAGAGTTCCACATCTTCTCTATATTGTATTTG |
All Microarray Probes Designed to Gene DMG400015488
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16082_PI426222305 | JHI_St_60k_v1 | DMT400040040 | GACCTGATAATGAAATCAGCTTGATCCAAAGAGTTCCACATCTTCTCTATATTGTATTTG |
Microarray Signals from CUST_16082_PI426222305
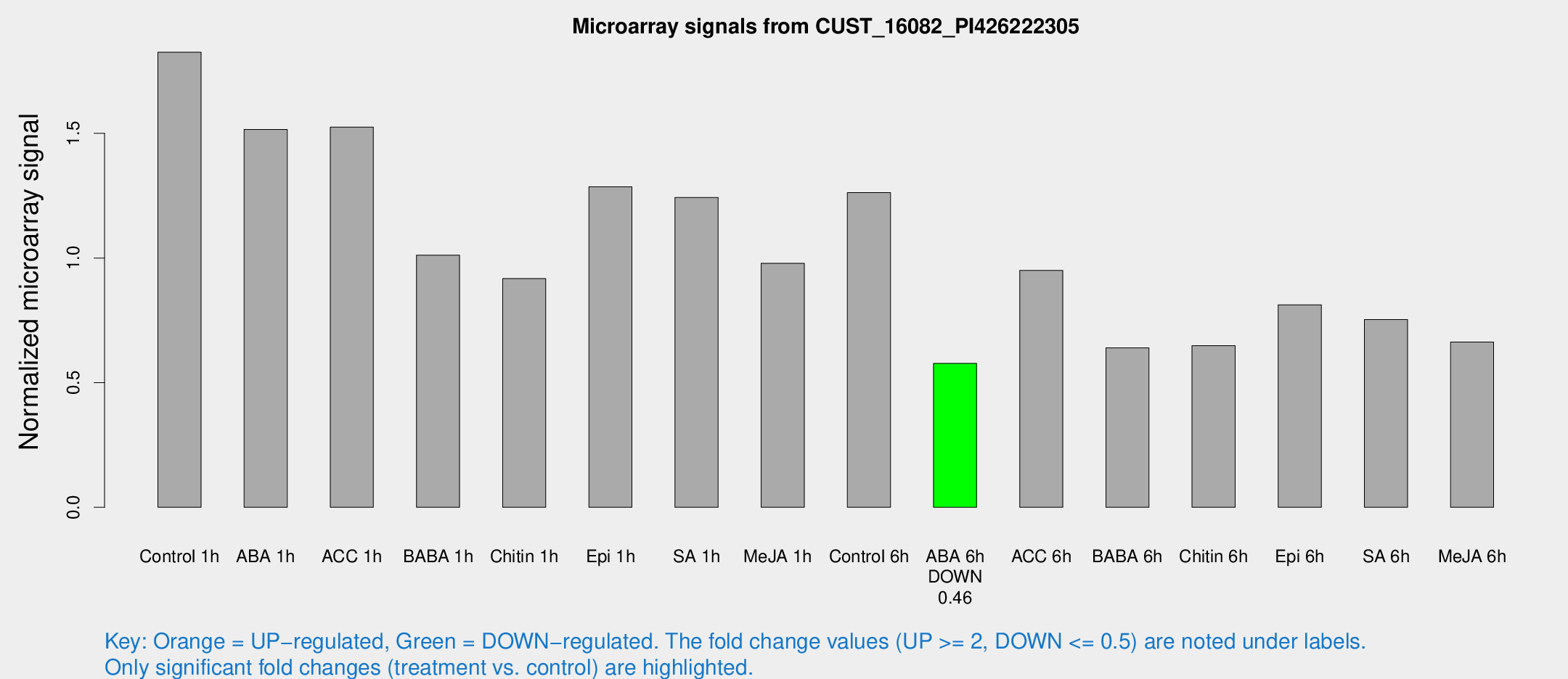
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11006.4 | 1012.27 | 1.8254 | 0.105391 |
| ABA 1h | 8078.19 | 694.549 | 1.5156 | 0.236588 |
| ACC 1h | 9487.45 | 1066.1 | 1.52511 | 0.0891915 |
| BABA 1h | 6055.02 | 1091.79 | 1.01105 | 0.0992878 |
| Chitin 1h | 4980.47 | 511.416 | 0.917202 | 0.130501 |
| Epi 1h | 6654.91 | 384.313 | 1.28591 | 0.0742448 |
| SA 1h | 7711.8 | 758.857 | 1.24275 | 0.0717526 |
| Me-JA 1h | 4786.7 | 276.993 | 0.978549 | 0.0565012 |
| Control 6h | 7693.55 | 980.71 | 1.2626 | 0.0910286 |
| ABA 6h | 3705.52 | 357.548 | 0.577248 | 0.0333326 |
| ACC 6h | 6573.3 | 628.564 | 0.950182 | 0.0548621 |
| BABA 6h | 4346.83 | 521.425 | 0.63981 | 0.0962275 |
| Chitin 6h | 4166.25 | 405.714 | 0.64874 | 0.0415899 |
| Epi 6h | 5478.74 | 316.486 | 0.812428 | 0.0653567 |
| SA 6h | 4733.02 | 1046.8 | 0.752819 | 0.107185 |
| Me-JA 6h | 3961.66 | 256.183 | 0.662746 | 0.0382684 |
Source Transcript PGSC0003DMT400040040 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G36530.2 | +2 | 3e-166 | 480 | 253/378 (67%) | alpha/beta-Hydrolases superfamily protein | chr4:17240120-17241770 REVERSE LENGTH=378 |
