Probe CUST_14842_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14842_PI426222305 | JHI_St_60k_v1 | DMT400065996 | CCAACGCAATTTTACTTAAGTGGATATTACATGATTGGAGCGACGGAGATTGTATAAAAA |
All Microarray Probes Designed to Gene DMG401025689
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14870_PI426222305 | JHI_St_60k_v1 | DMT400065993 | GTTTATGTTTGATTTATATGGCAAATAGTGGATATTACATGATTGGAGCGACGGAGATTG |
| CUST_14842_PI426222305 | JHI_St_60k_v1 | DMT400065996 | CCAACGCAATTTTACTTAAGTGGATATTACATGATTGGAGCGACGGAGATTGTATAAAAA |
Microarray Signals from CUST_14842_PI426222305
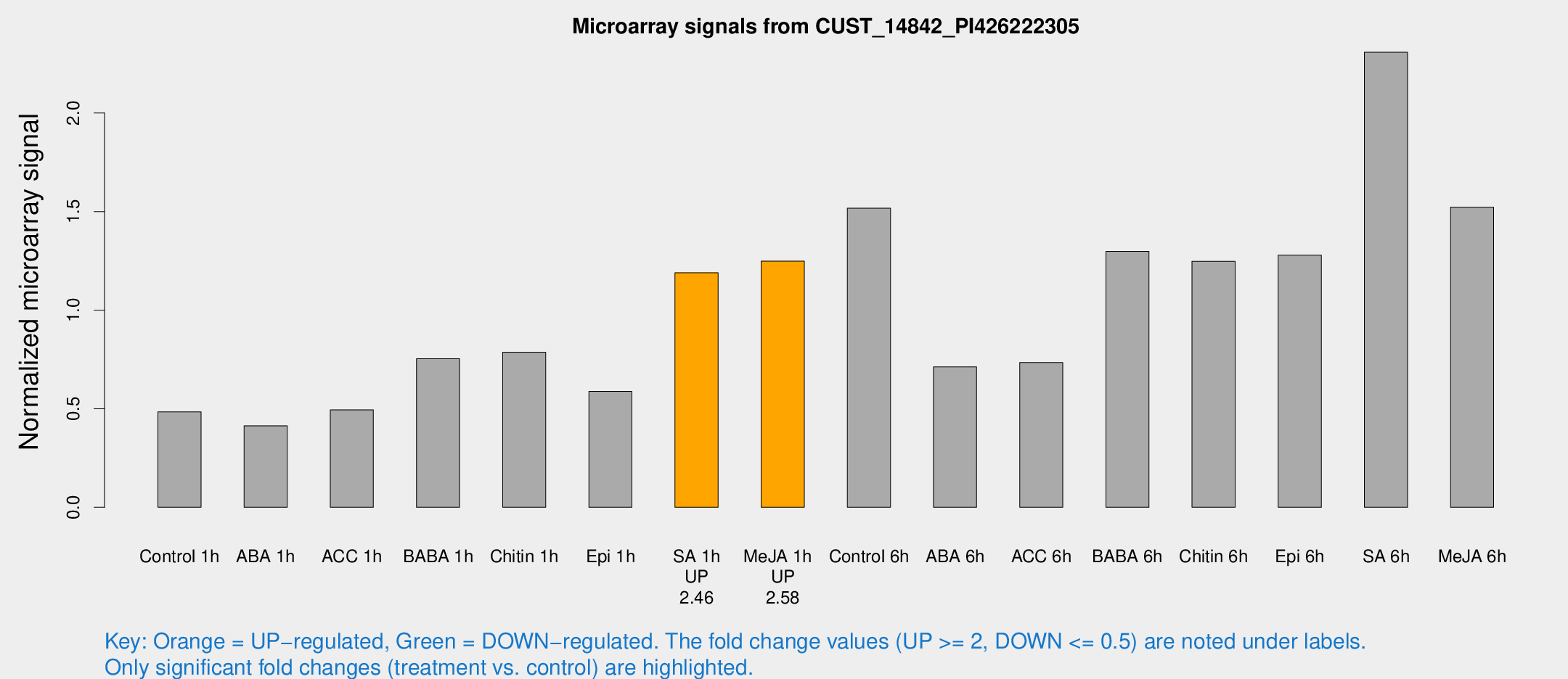
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 349.579 | 66.3082 | 0.484031 | 0.0573984 |
| ABA 1h | 279.63 | 85.739 | 0.41337 | 0.0985701 |
| ACC 1h | 363.431 | 54.931 | 0.494049 | 0.0450969 |
| BABA 1h | 528.518 | 99.3592 | 0.754181 | 0.0804793 |
| Chitin 1h | 500.088 | 57.9666 | 0.786498 | 0.045632 |
| Epi 1h | 371.566 | 76.1547 | 0.588353 | 0.114392 |
| SA 1h | 862.174 | 87.6474 | 1.18992 | 0.101671 |
| Me-JA 1h | 728.161 | 114.912 | 1.24841 | 0.107485 |
| Control 6h | 1125.33 | 243.547 | 1.51696 | 0.245926 |
| ABA 6h | 536.794 | 67.9822 | 0.712352 | 0.121396 |
| ACC 6h | 607.722 | 102.332 | 0.73411 | 0.116185 |
| BABA 6h | 1017.69 | 59.0023 | 1.29875 | 0.0751031 |
| Chitin 6h | 932.831 | 69.9749 | 1.24777 | 0.0721762 |
| Epi 6h | 1009.84 | 58.5887 | 1.27872 | 0.07395 |
| SA 6h | 1727.72 | 437.185 | 2.30843 | 0.428697 |
| Me-JA 6h | 1112.8 | 227.206 | 1.52238 | 0.236858 |
Source Transcript PGSC0003DMT400065996 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G35160.1 | +1 | 7e-63 | 212 | 137/382 (36%) | O-methyltransferase family protein | chr4:16730989-16732808 REVERSE LENGTH=382 |
