Probe CUST_14720_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14720_PI426222305 | JHI_St_60k_v1 | DMT400084751 | CCGATTGAACAAGAAGTAGAAAGTCCTATGCCTGCTAAGAAGCCGCGTTTATTTTTCGAT |
All Microarray Probes Designed to Gene DMG400034322
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14720_PI426222305 | JHI_St_60k_v1 | DMT400084751 | CCGATTGAACAAGAAGTAGAAAGTCCTATGCCTGCTAAGAAGCCGCGTTTATTTTTCGAT |
Microarray Signals from CUST_14720_PI426222305
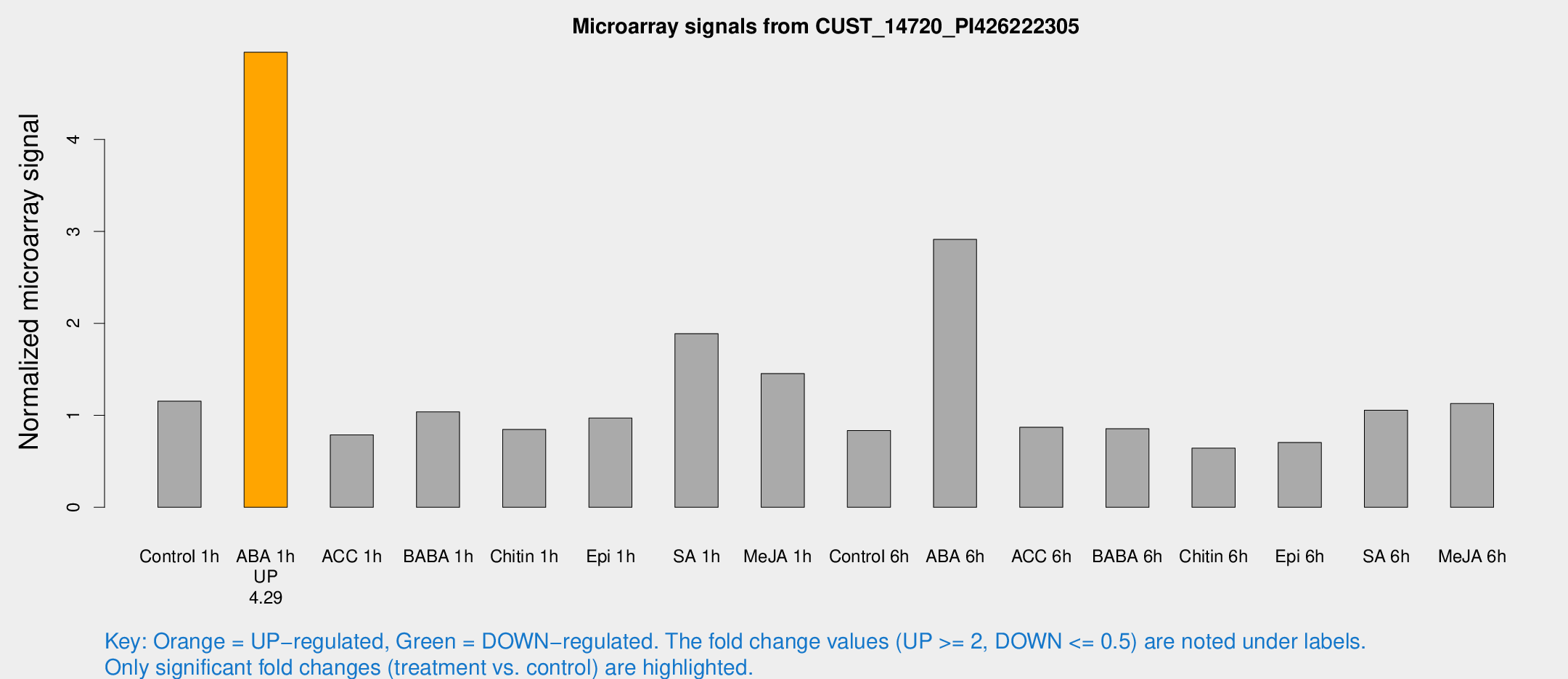
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 911.037 | 107.748 | 1.154 | 0.0667428 |
| ABA 1h | 3560.35 | 677.21 | 4.94818 | 0.805619 |
| ACC 1h | 721.673 | 226.256 | 0.786999 | 0.275359 |
| BABA 1h | 831.509 | 196.205 | 1.03782 | 0.173835 |
| Chitin 1h | 600.208 | 70.7224 | 0.845693 | 0.0490179 |
| Epi 1h | 655.173 | 38.0195 | 0.969925 | 0.0561785 |
| SA 1h | 1611.4 | 368.927 | 1.88833 | 0.405716 |
| Me-JA 1h | 960.097 | 189.224 | 1.45424 | 0.156127 |
| Control 6h | 666.625 | 94.2673 | 0.835285 | 0.0592076 |
| ABA 6h | 2634.26 | 676.307 | 2.91359 | 0.836522 |
| ACC 6h | 777.787 | 45.0713 | 0.870406 | 0.123133 |
| BABA 6h | 750.319 | 61.8057 | 0.854439 | 0.0494945 |
| Chitin 6h | 535.118 | 31.1883 | 0.643202 | 0.0417026 |
| Epi 6h | 630.714 | 80.7095 | 0.705227 | 0.0574987 |
| SA 6h | 828.339 | 111.52 | 1.05553 | 0.0668272 |
| Me-JA 6h | 888.922 | 115.487 | 1.128 | 0.0951343 |
Source Transcript PGSC0003DMT400084751 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G27730.1 | +1 | 2e-39 | 139 | 107/267 (40%) | salt tolerance zinc finger | chr1:9648302-9648985 REVERSE LENGTH=227 |
