Probe CUST_14717_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14717_PI426222305 | JHI_St_60k_v1 | DMT400066569 | CATAGACCAGTTCTATGATTGTTGCAAAAATATCATACCAAATGCTTATTGGTGGCATCA |
All Microarray Probes Designed to Gene DMG400025873
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14717_PI426222305 | JHI_St_60k_v1 | DMT400066569 | CATAGACCAGTTCTATGATTGTTGCAAAAATATCATACCAAATGCTTATTGGTGGCATCA |
Microarray Signals from CUST_14717_PI426222305
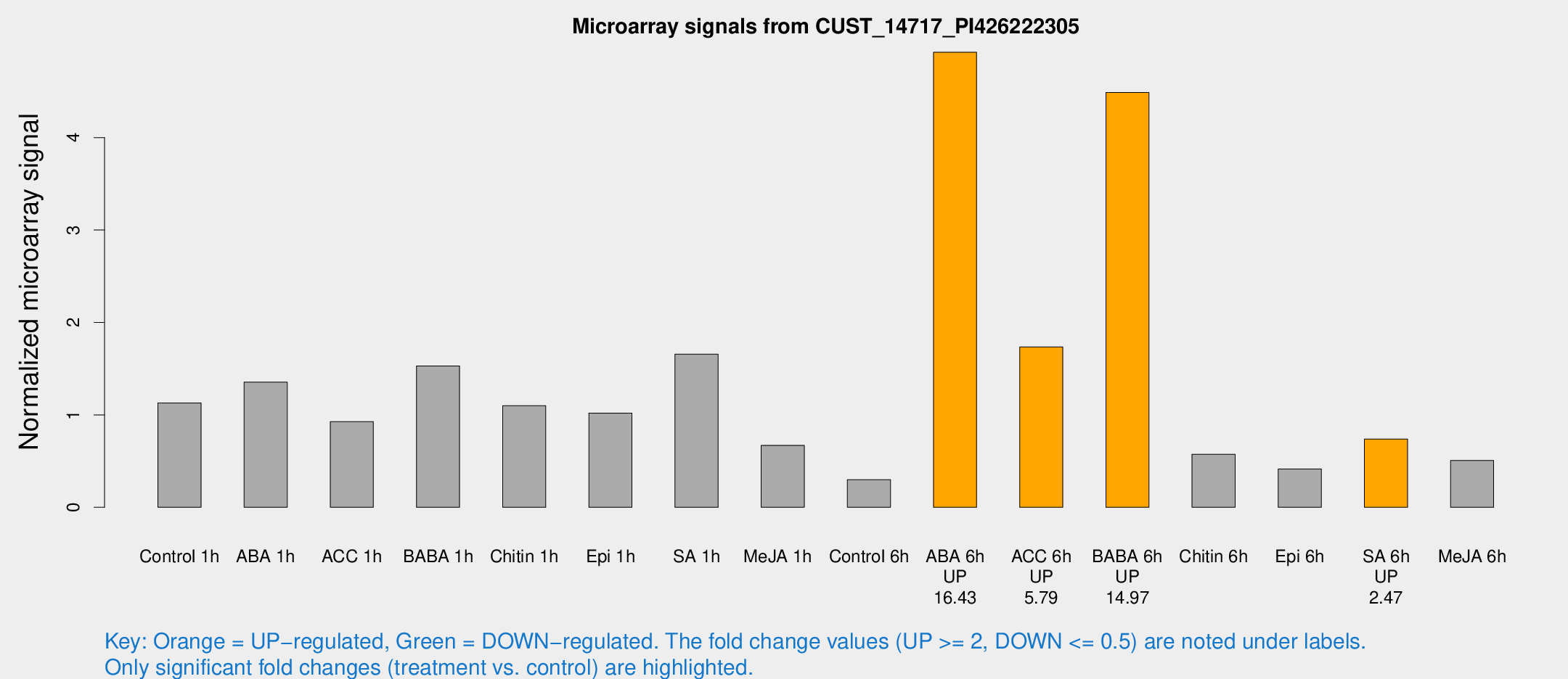
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 489.112 | 127.992 | 1.12909 | 0.217935 |
| ABA 1h | 573.767 | 191.981 | 1.35363 | 0.596464 |
| ACC 1h | 482.427 | 182.8 | 0.926135 | 0.478597 |
| BABA 1h | 691.453 | 214.874 | 1.52944 | 0.498263 |
| Chitin 1h | 449.867 | 151.431 | 1.09866 | 0.341487 |
| Epi 1h | 364.211 | 42.9824 | 1.01954 | 0.121541 |
| SA 1h | 786.375 | 293.579 | 1.65689 | 0.525083 |
| Me-JA 1h | 273.326 | 127.185 | 0.669969 | 0.285474 |
| Control 6h | 129.408 | 31.3405 | 0.299755 | 0.0570032 |
| ABA 6h | 2464.63 | 833.673 | 4.92373 | 1.90093 |
| ACC 6h | 812.93 | 47.2747 | 1.73591 | 0.159701 |
| BABA 6h | 2262.15 | 743.333 | 4.488 | 1.38483 |
| Chitin 6h | 275.821 | 85.6285 | 0.574296 | 0.191904 |
| Epi 6h | 240.941 | 113.154 | 0.414296 | 0.198603 |
| SA 6h | 300.35 | 29.1386 | 0.738977 | 0.102571 |
| Me-JA 6h | 248.25 | 107.774 | 0.506035 | 0.196856 |
Source Transcript PGSC0003DMT400066569 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15480.1 | +2 | 1e-146 | 437 | 225/487 (46%) | UDP-glucosyl transferase 73B5 | chr2:6758817-6760452 FORWARD LENGTH=484 |
