Probe CUST_14623_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14623_PI426222305 | JHI_St_60k_v1 | DMT400066574 | GCGTGTGTACGTTTTCTGTTTTGCTGATTTGATCATTACCTTTTGTCACCACTTCTAATA |
All Microarray Probes Designed to Gene DMG400025877
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14508_PI426222305 | JHI_St_60k_v1 | DMT400066573 | TCTTACAATGGATGGGCTACTTTGCTACAAGATATAACTTCATATACTTCCACAGGTCAT |
| CUST_14623_PI426222305 | JHI_St_60k_v1 | DMT400066574 | GCGTGTGTACGTTTTCTGTTTTGCTGATTTGATCATTACCTTTTGTCACCACTTCTAATA |
Microarray Signals from CUST_14623_PI426222305
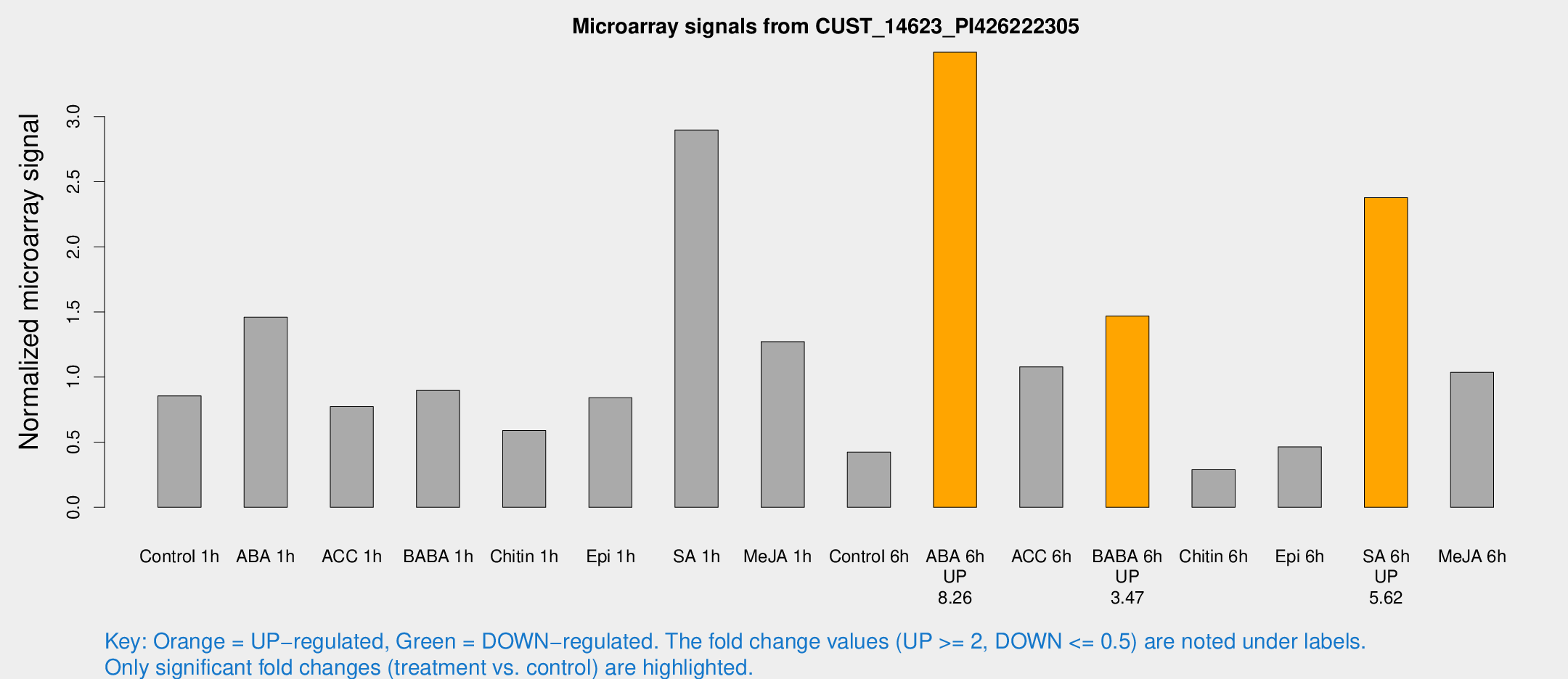
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2216.72 | 384.185 | 0.856017 | 0.106925 |
| ABA 1h | 3330.36 | 491.442 | 1.45983 | 0.178501 |
| ACC 1h | 2252.6 | 688.111 | 0.772696 | 0.243421 |
| BABA 1h | 2430.45 | 710.33 | 0.897779 | 0.232929 |
| Chitin 1h | 1386.74 | 277.119 | 0.589717 | 0.090337 |
| Epi 1h | 1855.12 | 205.971 | 0.841476 | 0.0976998 |
| SA 1h | 8115.42 | 2134.52 | 2.89657 | 0.699051 |
| Me-JA 1h | 2738.95 | 587.921 | 1.27116 | 0.169238 |
| Control 6h | 1093.46 | 162.221 | 0.423089 | 0.0344771 |
| ABA 6h | 10589.4 | 3353.39 | 3.49429 | 1.20118 |
| ACC 6h | 3213.88 | 592.353 | 1.07838 | 0.309862 |
| BABA 6h | 4299.98 | 778.91 | 1.46756 | 0.332859 |
| Chitin 6h | 781.811 | 77.7445 | 0.288829 | 0.0255358 |
| Epi 6h | 1385.58 | 312.511 | 0.464031 | 0.0706218 |
| SA 6h | 6085.33 | 1038.79 | 2.37684 | 0.278466 |
| Me-JA 6h | 2797.86 | 667.564 | 1.03688 | 0.344913 |
Source Transcript PGSC0003DMT400066574 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34131.1 | +1 | 7e-178 | 514 | 261/478 (55%) | UDP-glucosyl transferase 73B3 | chr4:16343268-16344713 REVERSE LENGTH=481 |
