Probe CUST_1459_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_1459_PI426222305 | JHI_St_60k_v1 | DMT400026191 | CCTTGAAACTTCTCTCCCGTACCTCAAAGGTATTTTTTCACACAATCAATAATCTATAAC |
All Microarray Probes Designed to Gene DMG400010092
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_1459_PI426222305 | JHI_St_60k_v1 | DMT400026191 | CCTTGAAACTTCTCTCCCGTACCTCAAAGGTATTTTTTCACACAATCAATAATCTATAAC |
| CUST_1545_PI426222305 | JHI_St_60k_v1 | DMT400026190 | CATATAATATGATCATTTACTCTCACTAACTCTAGATTCTGGATTCACCTCGTCTCTACT |
| CUST_895_PI426222305 | JHI_St_60k_v1 | DMT400026192 | CACAACGCCTTCCTTGCAAACACTGAATAGTAATATTTGTATGGTACTTACCTCAAAATA |
Microarray Signals from CUST_1459_PI426222305
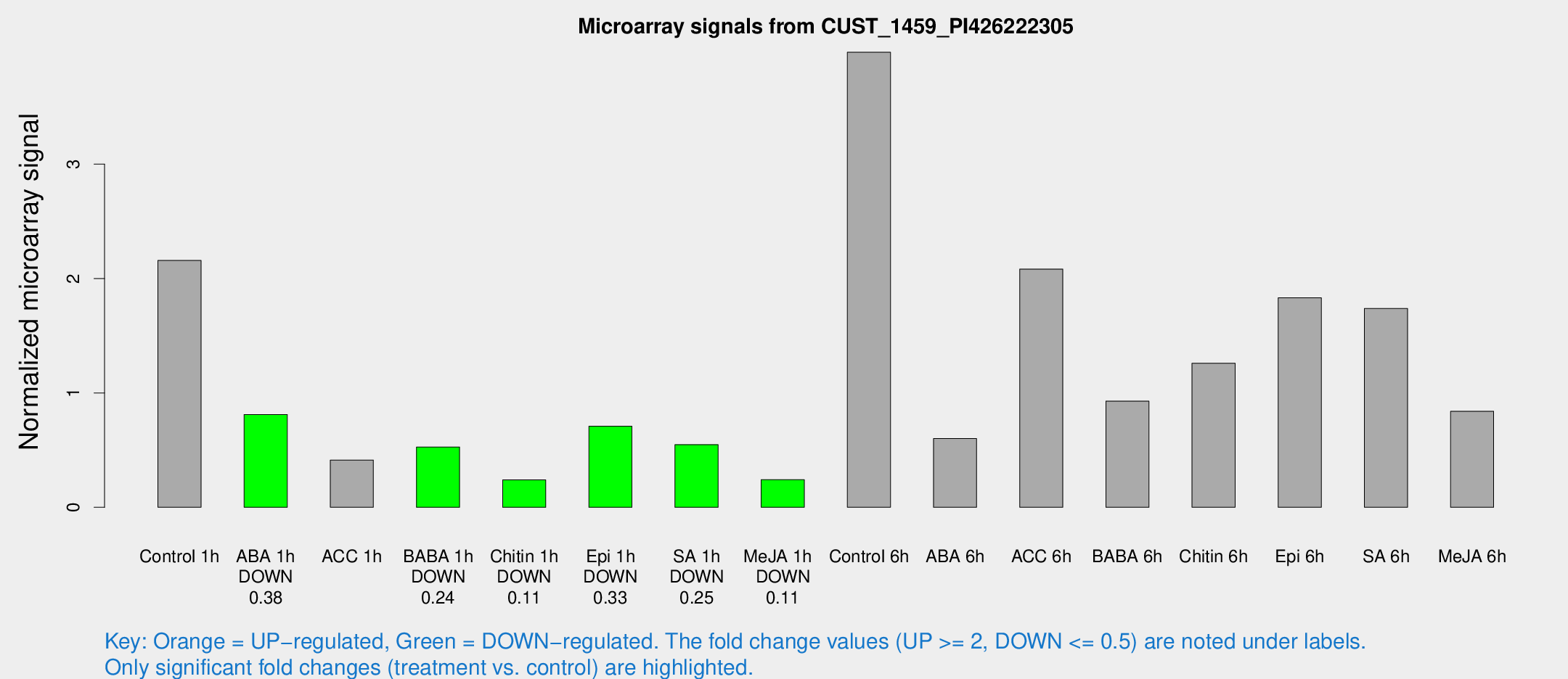
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1382.85 | 256.856 | 2.1585 | 0.320561 |
| ABA 1h | 454.998 | 64.7537 | 0.811581 | 0.179364 |
| ACC 1h | 478.888 | 220.992 | 0.413278 | 0.765209 |
| BABA 1h | 332.146 | 72.228 | 0.527094 | 0.0748517 |
| Chitin 1h | 135.299 | 15.9089 | 0.239203 | 0.0323002 |
| Epi 1h | 393.14 | 72.0988 | 0.708871 | 0.13206 |
| SA 1h | 368.401 | 87.8473 | 0.547672 | 0.150359 |
| Me-JA 1h | 122.967 | 8.47312 | 0.241853 | 0.0383347 |
| Control 6h | 2748.42 | 923.801 | 3.97914 | 1.39983 |
| ABA 6h | 422.749 | 102.519 | 0.600999 | 0.195129 |
| ACC 6h | 1548.33 | 299.952 | 2.08298 | 0.419688 |
| BABA 6h | 712.303 | 228.797 | 0.928248 | 0.326988 |
| Chitin 6h | 833.817 | 48.4434 | 1.26031 | 0.0730124 |
| Epi 6h | 1297.14 | 146.493 | 1.83172 | 0.105918 |
| SA 6h | 1356.18 | 677.372 | 1.73879 | 0.793644 |
| Me-JA 6h | 535.138 | 89.4546 | 0.839618 | 0.19294 |
Source Transcript PGSC0003DMT400026191 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc02g089770.2 | +3 | 3e-163 | 471 | 229/240 (95%) | genomic_reference:SL2.50ch02 gene_region:46055433-46058696 transcript_region:SL2.50ch02:46055433..46058696+ go_terms:GO:0004022,GO:0045551 functional_description:Dihydroflavonol-4-reductase (AHRD V1 ***- B6TK03_MAIZE); contains Interpro domain(s) IPR016040 NAD(P)-binding domain |
| TAIR PP10 | AT5G14700.1 | +3 | 8e-58 | 199 | 98/213 (46%) | NAD(P)-binding Rossmann-fold superfamily protein | chr5:4740502-4743327 REVERSE LENGTH=368 |
