Probe CUST_14598_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14598_PI426222305 | JHI_St_60k_v1 | DMT400066516 | TGGTTCAATTTCCTGTCAACGTGCTTTCCAAAGCGTGATGTCATAACAACATTTAAAAAA |
All Microarray Probes Designed to Gene DMG400025847
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14598_PI426222305 | JHI_St_60k_v1 | DMT400066516 | TGGTTCAATTTCCTGTCAACGTGCTTTCCAAAGCGTGATGTCATAACAACATTTAAAAAA |
| CUST_14396_PI426222305 | JHI_St_60k_v1 | DMT400066515 | TAATCCTTGTATTGGTTGCTACCTTAGGTGAAAGTAGTTCAGAGATTGTTGCGCGACTCA |
Microarray Signals from CUST_14598_PI426222305
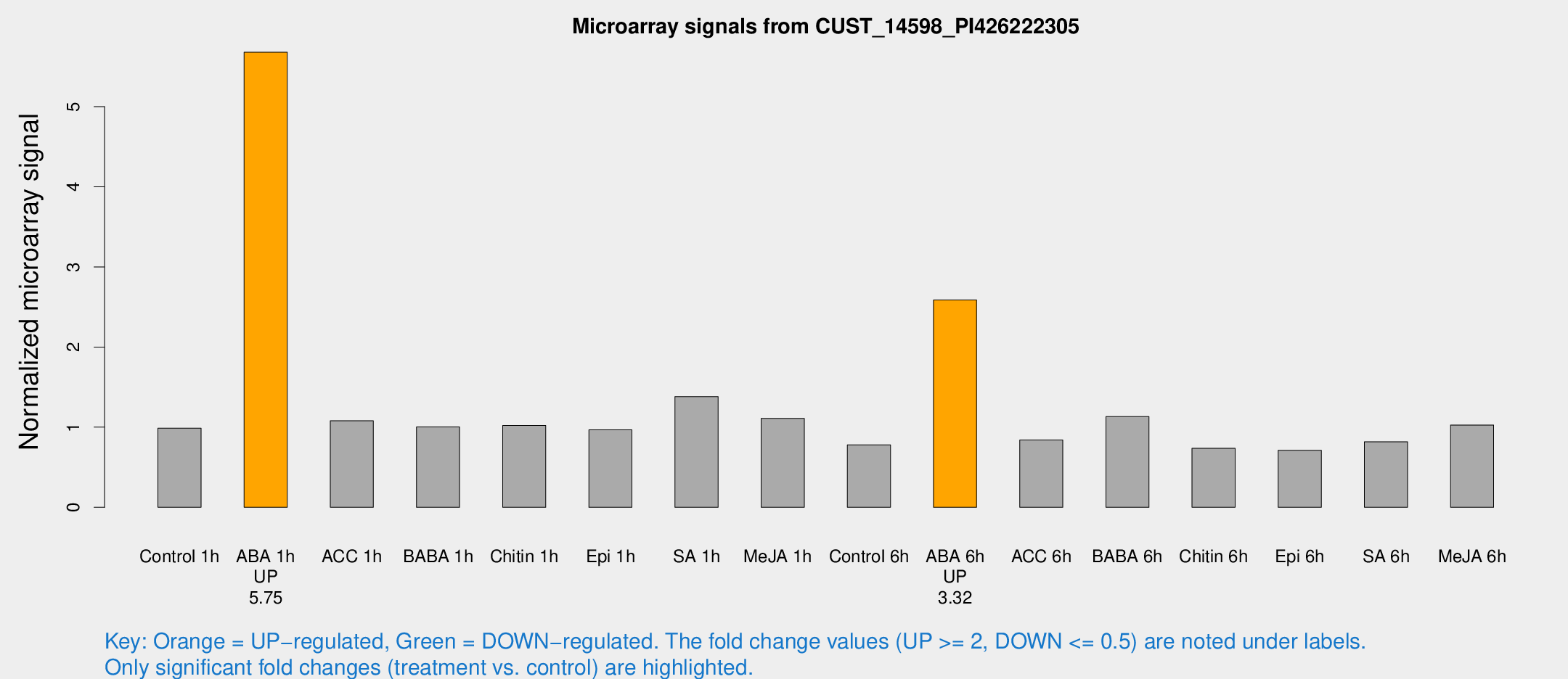
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 204.655 | 26.5932 | 0.987672 | 0.0637034 |
| ABA 1h | 1031.67 | 87.0394 | 5.67962 | 0.328284 |
| ACC 1h | 245.915 | 62.1614 | 1.07928 | 0.259045 |
| BABA 1h | 209.328 | 45.9766 | 1.00316 | 0.153694 |
| Chitin 1h | 191.119 | 27.815 | 1.02128 | 0.066852 |
| Epi 1h | 170.747 | 10.2736 | 0.966586 | 0.0581334 |
| SA 1h | 304.392 | 61.9768 | 1.3811 | 0.258709 |
| Me-JA 1h | 187.28 | 20.9524 | 1.11011 | 0.0665602 |
| Control 6h | 170.868 | 44.6724 | 0.780153 | 0.161966 |
| ABA 6h | 574.369 | 82.3109 | 2.5883 | 0.306199 |
| ACC 6h | 200.277 | 28.589 | 0.83995 | 0.0506802 |
| BABA 6h | 260.322 | 22.5731 | 1.13214 | 0.0885119 |
| Chitin 6h | 160.442 | 9.86377 | 0.736903 | 0.0506269 |
| Epi 6h | 167.211 | 23.9627 | 0.71049 | 0.0681132 |
| SA 6h | 187.866 | 61.4079 | 0.817234 | 0.231133 |
| Me-JA 6h | 211.557 | 25.1412 | 1.02684 | 0.0611406 |
Source Transcript PGSC0003DMT400066516 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G33905.1 | +2 | 3e-87 | 270 | 130/216 (60%) | Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein | chr4:16254065-16255592 REVERSE LENGTH=261 |
