Probe CUST_14578_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14578_PI426222305 | JHI_St_60k_v1 | DMT400066555 | TTGAAATCACAGAAATCTACGTCGATTTGTCAGCAAGCTTATGGTGGGAAACCAAAGTCT |
All Microarray Probes Designed to Gene DMG400025867
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14714_PI426222305 | JHI_St_60k_v1 | DMT400066556 | CAAGCTTATGGTGGGAAACCAAAGTCTTAGTTGTGTCAGTCAATGTTTCTGAGATTTTTT |
| CUST_14578_PI426222305 | JHI_St_60k_v1 | DMT400066555 | TTGAAATCACAGAAATCTACGTCGATTTGTCAGCAAGCTTATGGTGGGAAACCAAAGTCT |
Microarray Signals from CUST_14578_PI426222305
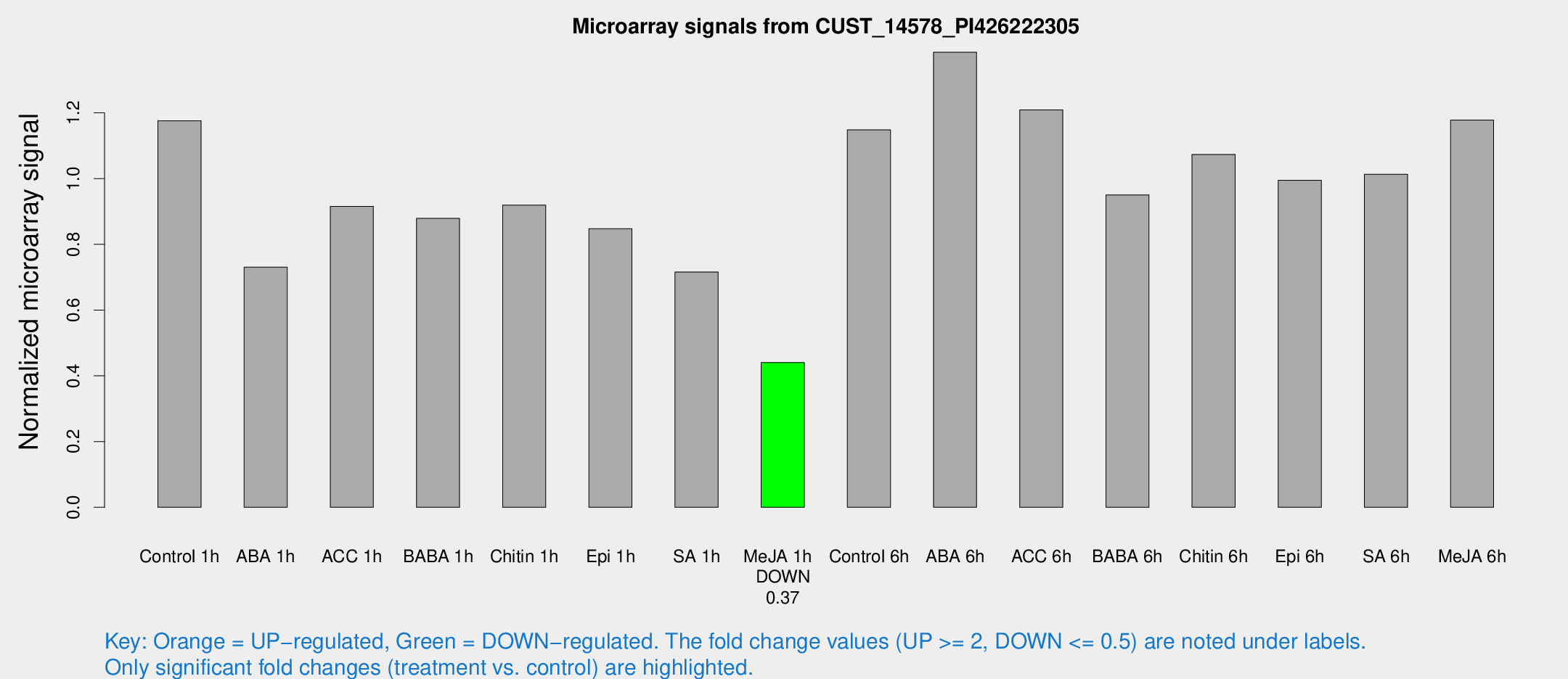
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2954.5 | 364.793 | 1.17585 | 0.0712506 |
| ABA 1h | 1603.95 | 92.91 | 0.730905 | 0.0422188 |
| ACC 1h | 2449.13 | 508.681 | 0.915295 | 0.147454 |
| BABA 1h | 2196.77 | 453.841 | 0.879046 | 0.112525 |
| Chitin 1h | 2077.86 | 249.082 | 0.919591 | 0.0585658 |
| Epi 1h | 1825.82 | 141.694 | 0.847426 | 0.0545418 |
| SA 1h | 1852.46 | 240.288 | 0.716031 | 0.0546705 |
| Me-JA 1h | 893.859 | 62.3045 | 0.440412 | 0.0271267 |
| Control 6h | 2985.31 | 613.756 | 1.1482 | 0.168112 |
| ABA 6h | 3641.64 | 210.323 | 1.38447 | 0.0799416 |
| ACC 6h | 3461.82 | 328.728 | 1.20926 | 0.0698277 |
| BABA 6h | 2643.92 | 176.656 | 0.950527 | 0.0548924 |
| Chitin 6h | 2830.06 | 163.656 | 1.07332 | 0.0653806 |
| Epi 6h | 2797.08 | 250.287 | 0.994901 | 0.0769083 |
| SA 6h | 2604.03 | 524.565 | 1.01327 | 0.114236 |
| Me-JA 6h | 2948.92 | 376.5 | 1.17797 | 0.0680217 |
Source Transcript PGSC0003DMT400066555 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15320.1 | +1 | 3e-82 | 253 | 124/215 (58%) | Leucine-rich repeat (LRR) family protein | chr2:6666527-6667675 REVERSE LENGTH=382 |
