Probe CUST_14571_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14571_PI426222305 | JHI_St_60k_v1 | DMT400066576 | TTATATGTGGTGCGAGGTCAATTGAGCGAACTAACTTTGAACCACACGTCCGTTATACCG |
All Microarray Probes Designed to Gene DMG400025878
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14464_PI426222305 | JHI_St_60k_v1 | DMT400066575 | TTATATGTGGTGCGAGGTCAATTGAGCGAACTAACTTTGAACCACACGTCCGTTATACCG |
| CUST_14571_PI426222305 | JHI_St_60k_v1 | DMT400066576 | TTATATGTGGTGCGAGGTCAATTGAGCGAACTAACTTTGAACCACACGTCCGTTATACCG |
Microarray Signals from CUST_14571_PI426222305
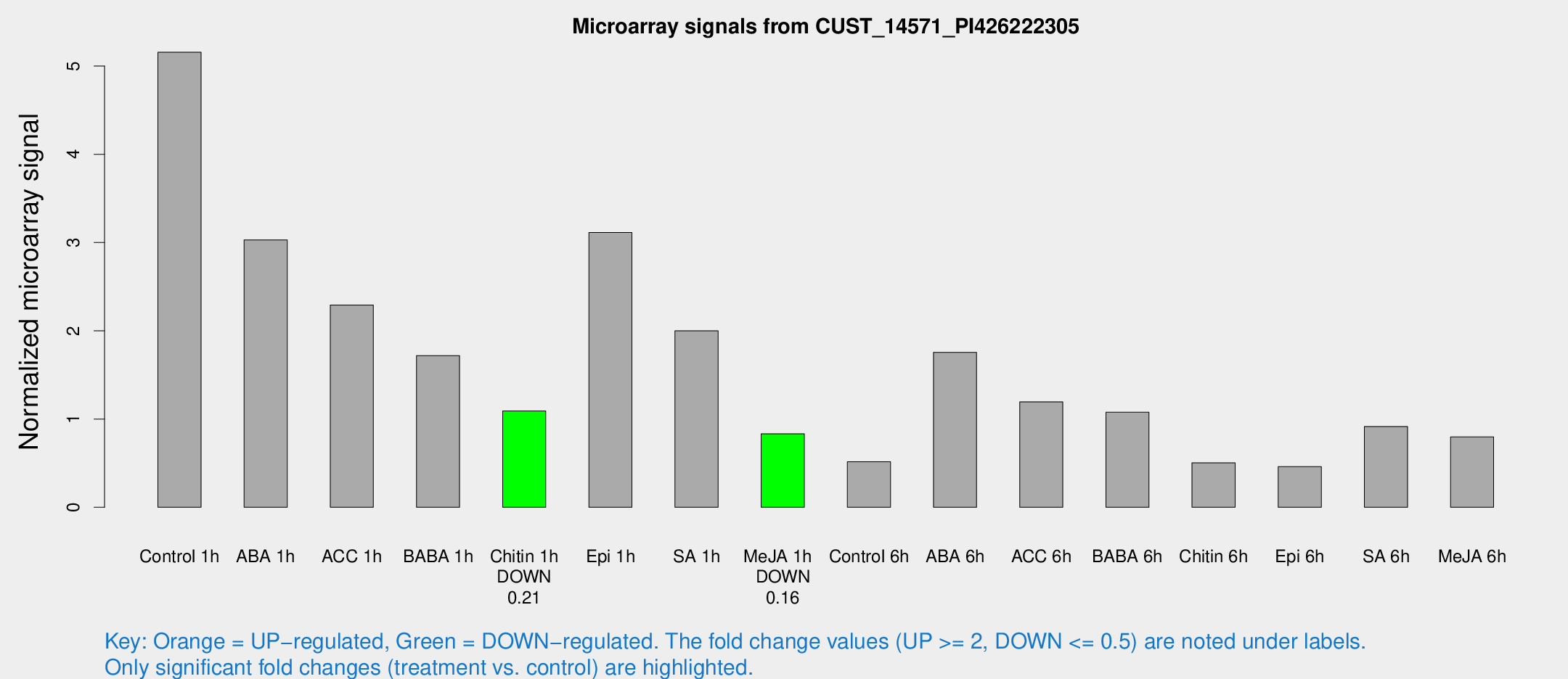
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 798.573 | 77.0601 | 5.15597 | 0.453811 |
| ABA 1h | 571.559 | 326.04 | 3.03028 | 2.14945 |
| ACC 1h | 592.866 | 323.423 | 2.29174 | 2.67039 |
| BABA 1h | 321.109 | 130.99 | 1.71905 | 0.839377 |
| Chitin 1h | 162.034 | 46.2383 | 1.09115 | 0.220893 |
| Epi 1h | 438.523 | 99.8958 | 3.11335 | 0.900276 |
| SA 1h | 355.543 | 126.454 | 1.99952 | 0.596755 |
| Me-JA 1h | 107.235 | 17.6194 | 0.831916 | 0.0717173 |
| Control 6h | 85.0653 | 21.4931 | 0.516877 | 0.103182 |
| ABA 6h | 330.789 | 102.954 | 1.75617 | 0.71815 |
| ACC 6h | 211.369 | 15.3781 | 1.19465 | 0.225095 |
| BABA 6h | 189.422 | 30.0461 | 1.07729 | 0.13781 |
| Chitin 6h | 84.0237 | 11.9856 | 0.503508 | 0.0705228 |
| Epi 6h | 92.353 | 29.2084 | 0.460375 | 0.179569 |
| SA 6h | 148.884 | 35.3186 | 0.915905 | 0.148923 |
| Me-JA 6h | 123.123 | 12.268 | 0.797461 | 0.0953753 |
Source Transcript PGSC0003DMT400066576 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15480.1 | +3 | 5e-172 | 502 | 263/475 (55%) | UDP-glucosyl transferase 73B5 | chr2:6758817-6760452 FORWARD LENGTH=484 |
