Probe CUST_14544_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14544_PI426222305 | JHI_St_60k_v1 | DMT400066567 | CATAGACCAGTTCTATGATTGTTGCAAAAATATCATACCAAATGCTTATTGGTGGCATCA |
All Microarray Probes Designed to Gene DMG400025871
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14544_PI426222305 | JHI_St_60k_v1 | DMT400066567 | CATAGACCAGTTCTATGATTGTTGCAAAAATATCATACCAAATGCTTATTGGTGGCATCA |
Microarray Signals from CUST_14544_PI426222305
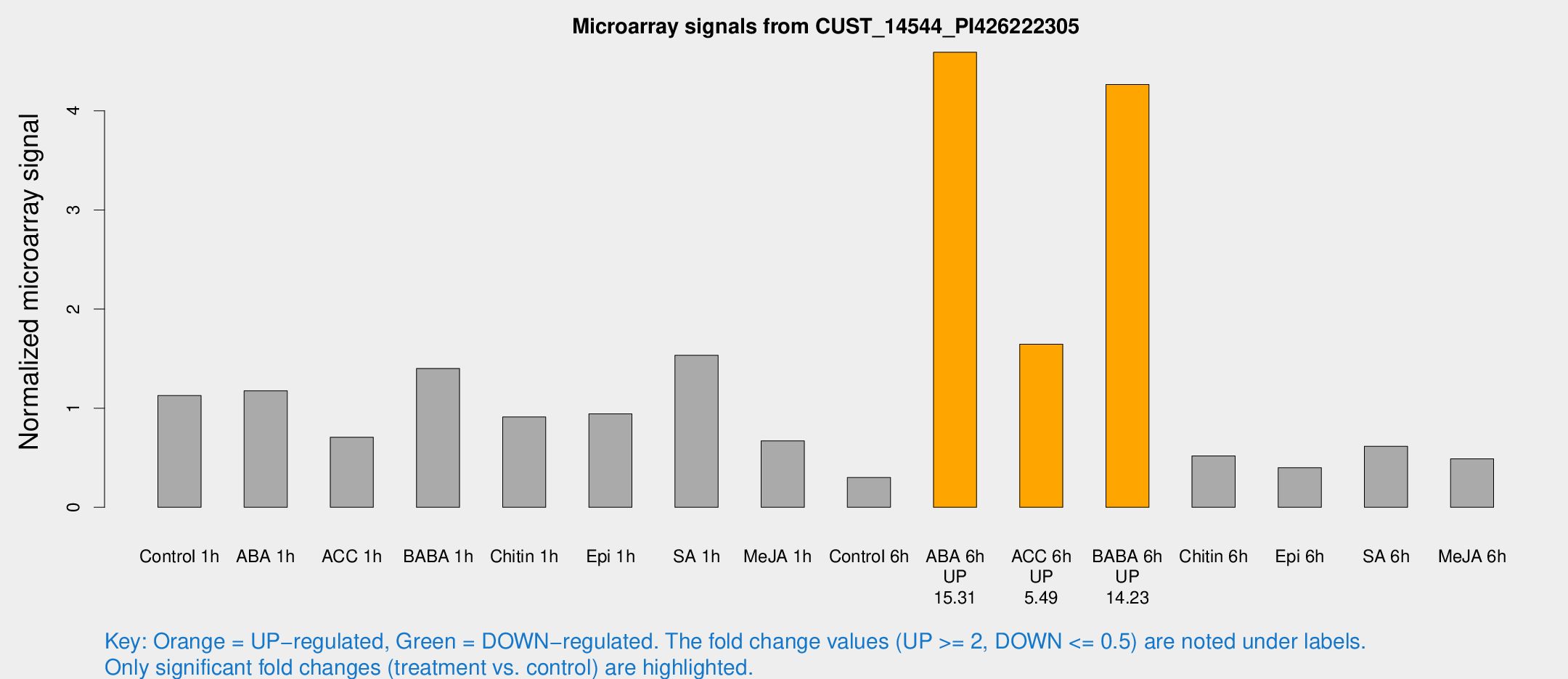
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 547.067 | 121.9 | 1.12714 | 0.183227 |
| ABA 1h | 578.87 | 200.829 | 1.17531 | 0.567502 |
| ACC 1h | 493.557 | 217.786 | 0.707559 | 0.645854 |
| BABA 1h | 724.588 | 229.563 | 1.40078 | 0.467688 |
| Chitin 1h | 416.65 | 121.848 | 0.910942 | 0.273922 |
| Epi 1h | 380.748 | 30.6858 | 0.942058 | 0.0679492 |
| SA 1h | 850.177 | 348.038 | 1.5334 | 0.539609 |
| Me-JA 1h | 298.674 | 125.726 | 0.670527 | 0.236219 |
| Control 6h | 147.831 | 34.5225 | 0.299812 | 0.0535107 |
| ABA 6h | 2591.39 | 837.772 | 4.59133 | 1.69905 |
| ACC 6h | 886.948 | 99.3479 | 1.64597 | 0.154215 |
| BABA 6h | 2399.45 | 710.829 | 4.26494 | 1.13261 |
| Chitin 6h | 284.198 | 87.0289 | 0.517927 | 0.180914 |
| Epi 6h | 252.491 | 109.987 | 0.399128 | 0.159196 |
| SA 6h | 286.546 | 32.3484 | 0.616147 | 0.121705 |
| Me-JA 6h | 259.046 | 95.8444 | 0.488987 | 0.149437 |
Source Transcript PGSC0003DMT400066567 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15480.1 | +1 | 2e-146 | 437 | 225/487 (46%) | UDP-glucosyl transferase 73B5 | chr2:6758817-6760452 FORWARD LENGTH=484 |
