Probe CUST_14399_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14399_PI426222305 | JHI_St_60k_v1 | DMT400066736 | GTTGTAAATTCTAAGTAACCATGTGTTTGAATCATCCAATGGAACGTCTTTGAATGTACC |
All Microarray Probes Designed to Gene DMG400025932
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14773_PI426222305 | JHI_St_60k_v1 | DMT400066737 | GTTGTAAATTCTAAGTAACCATGTGTTTGAATCATCCAATGGAACGTCTTTGAATGTACC |
| CUST_14399_PI426222305 | JHI_St_60k_v1 | DMT400066736 | GTTGTAAATTCTAAGTAACCATGTGTTTGAATCATCCAATGGAACGTCTTTGAATGTACC |
Microarray Signals from CUST_14399_PI426222305
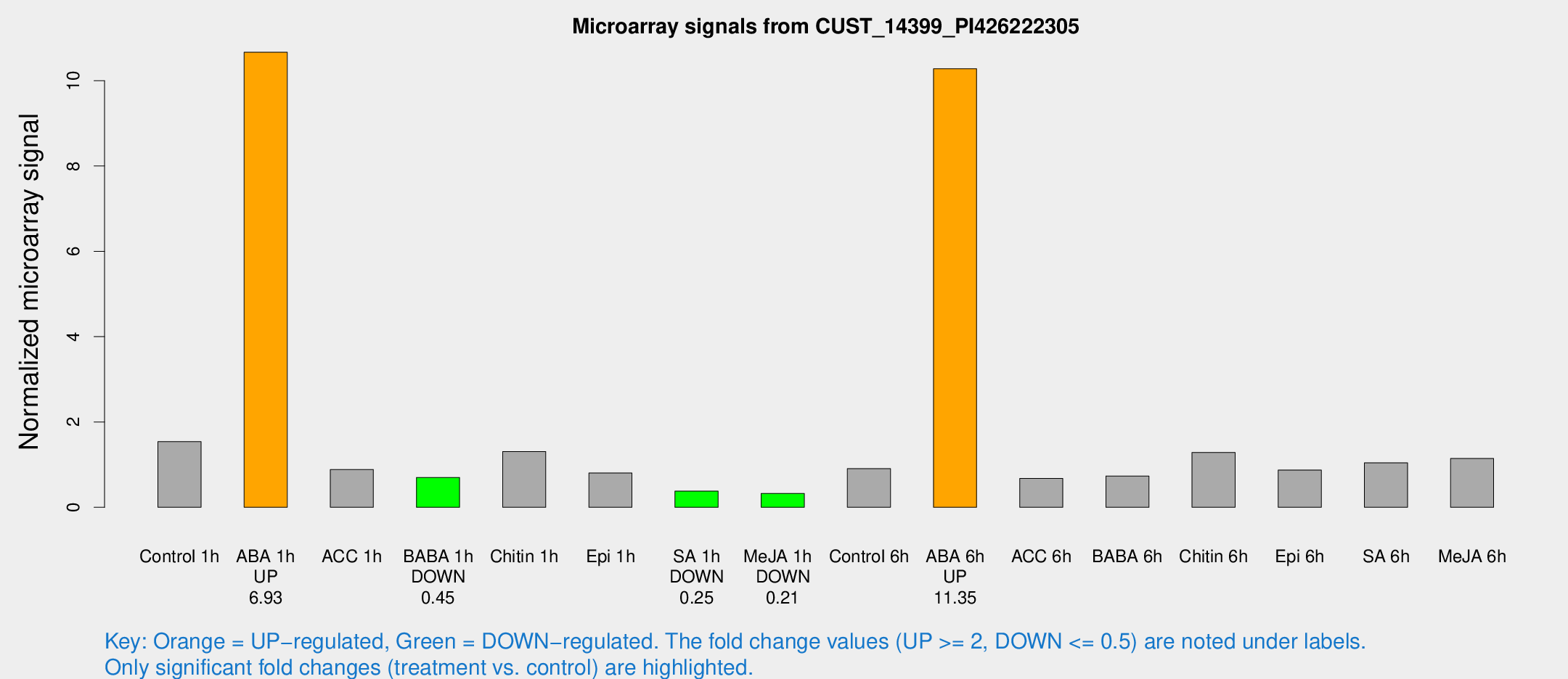
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 127.896 | 9.71933 | 1.53933 | 0.0972102 |
| ABA 1h | 791.104 | 95.9809 | 10.6669 | 1.85322 |
| ACC 1h | 83.9643 | 28.8709 | 0.886004 | 0.24991 |
| BABA 1h | 57.9155 | 10.4635 | 0.699707 | 0.0691385 |
| Chitin 1h | 102.148 | 23.8827 | 1.30847 | 0.201293 |
| Epi 1h | 57.8016 | 4.61824 | 0.803492 | 0.064551 |
| SA 1h | 35.7371 | 12.0408 | 0.378616 | 0.112669 |
| Me-JA 1h | 22.3342 | 3.56403 | 0.325103 | 0.0536583 |
| Control 6h | 80.4807 | 19.0482 | 0.905803 | 0.175751 |
| ABA 6h | 938.39 | 170.391 | 10.2792 | 1.63503 |
| ACC 6h | 68.1229 | 17.7006 | 0.675775 | 0.219919 |
| BABA 6h | 74.2754 | 19.3209 | 0.733575 | 0.285469 |
| Chitin 6h | 117.185 | 23.2129 | 1.28555 | 0.283247 |
| Epi 6h | 86.9778 | 22.7475 | 0.874293 | 0.200847 |
| SA 6h | 85.3851 | 6.08329 | 1.04085 | 0.133222 |
| Me-JA 6h | 104.714 | 33.096 | 1.14301 | 0.386608 |
Source Transcript PGSC0003DMT400066736 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34380.1 | +2 | 2e-123 | 318 | 159/243 (65%) | Transducin/WD40 repeat-like superfamily protein | chr4:16438835-16440322 FORWARD LENGTH=495 |
