Probe CUST_14396_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14396_PI426222305 | JHI_St_60k_v1 | DMT400066515 | TAATCCTTGTATTGGTTGCTACCTTAGGTGAAAGTAGTTCAGAGATTGTTGCGCGACTCA |
All Microarray Probes Designed to Gene DMG400025847
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14598_PI426222305 | JHI_St_60k_v1 | DMT400066516 | TGGTTCAATTTCCTGTCAACGTGCTTTCCAAAGCGTGATGTCATAACAACATTTAAAAAA |
| CUST_14396_PI426222305 | JHI_St_60k_v1 | DMT400066515 | TAATCCTTGTATTGGTTGCTACCTTAGGTGAAAGTAGTTCAGAGATTGTTGCGCGACTCA |
Microarray Signals from CUST_14396_PI426222305
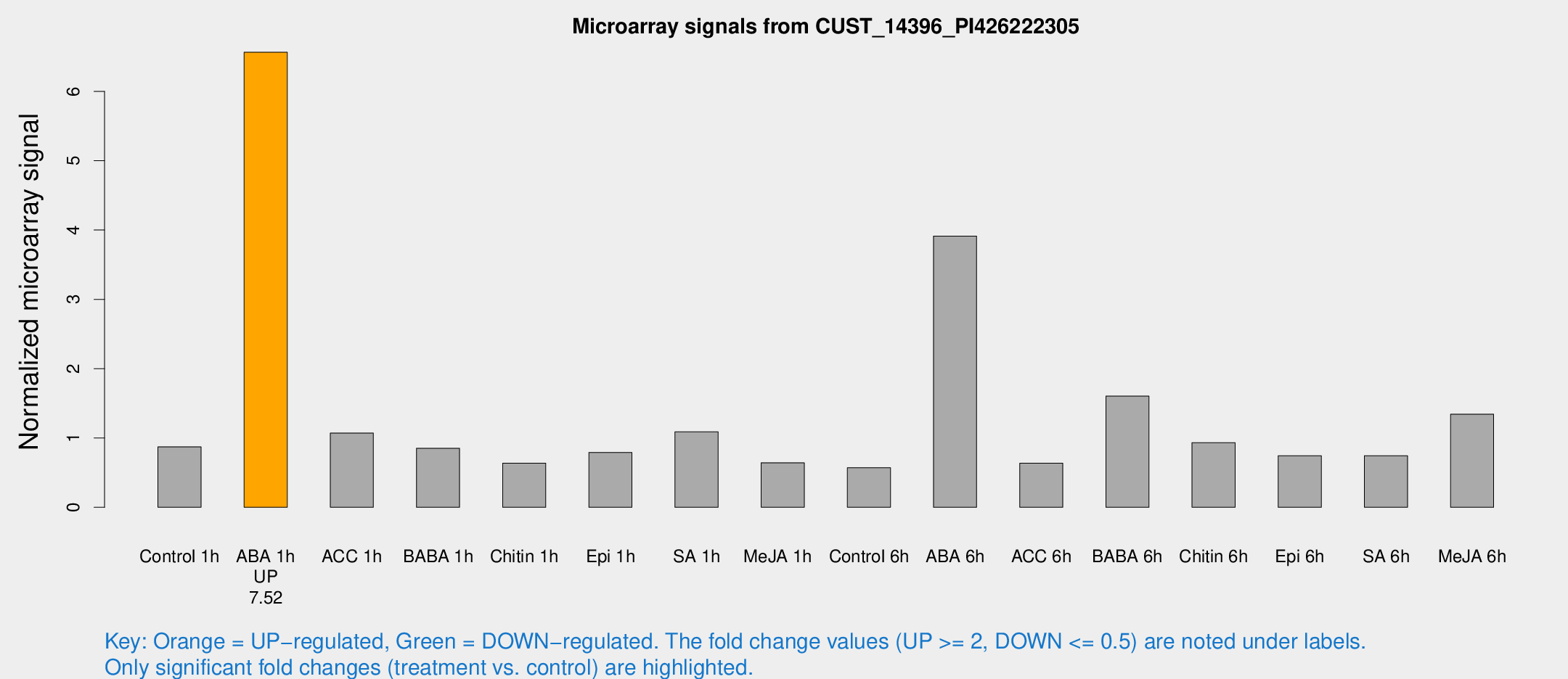
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13.5624 | 5.27828 | 0.872829 | 0.403233 |
| ABA 1h | 76.4007 | 6.47158 | 6.56531 | 0.454056 |
| ACC 1h | 14.6508 | 3.459 | 1.07174 | 0.259018 |
| BABA 1h | 12.575 | 5.16692 | 0.851955 | 0.359629 |
| Chitin 1h | 8.73386 | 3.96494 | 0.63767 | 0.285465 |
| Epi 1h | 8.94772 | 2.95335 | 0.791651 | 0.26124 |
| SA 1h | 15.3423 | 3.45987 | 1.08801 | 0.243667 |
| Me-JA 1h | 6.99125 | 3.04882 | 0.642958 | 0.294949 |
| Control 6h | 8.60299 | 3.38222 | 0.571694 | 0.267677 |
| ABA 6h | 57.8476 | 13.2981 | 3.91355 | 0.84703 |
| ACC 6h | 10.5844 | 3.78216 | 0.635904 | 0.257183 |
| BABA 6h | 23.7941 | 3.67569 | 1.60393 | 0.251813 |
| Chitin 6h | 13.2622 | 3.44894 | 0.932284 | 0.259005 |
| Epi 6h | 11.4536 | 3.59191 | 0.744456 | 0.255123 |
| SA 6h | 11.7694 | 5.49509 | 0.745136 | 0.31317 |
| Me-JA 6h | 17.8501 | 3.21423 | 1.34328 | 0.254951 |
Source Transcript PGSC0003DMT400066515 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G33905.1 | +2 | 1e-28 | 110 | 47/66 (71%) | Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein | chr4:16254065-16255592 REVERSE LENGTH=261 |
