Probe CUST_14384_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14384_PI426222305 | JHI_St_60k_v1 | DMT400059947 | AAAGCTCATGATGTCTATGGCTACCATGTTGATGGACCATTAAAGCTACCTTGCTCCGTC |
All Microarray Probes Designed to Gene DMG400023325
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14384_PI426222305 | JHI_St_60k_v1 | DMT400059947 | AAAGCTCATGATGTCTATGGCTACCATGTTGATGGACCATTAAAGCTACCTTGCTCCGTC |
| CUST_14276_PI426222305 | JHI_St_60k_v1 | DMT400059948 | AAACAATCACAACCATCATCACCACCAATATCTTTCTAGTTCTTTGTCTTTTCACTCATG |
Microarray Signals from CUST_14384_PI426222305
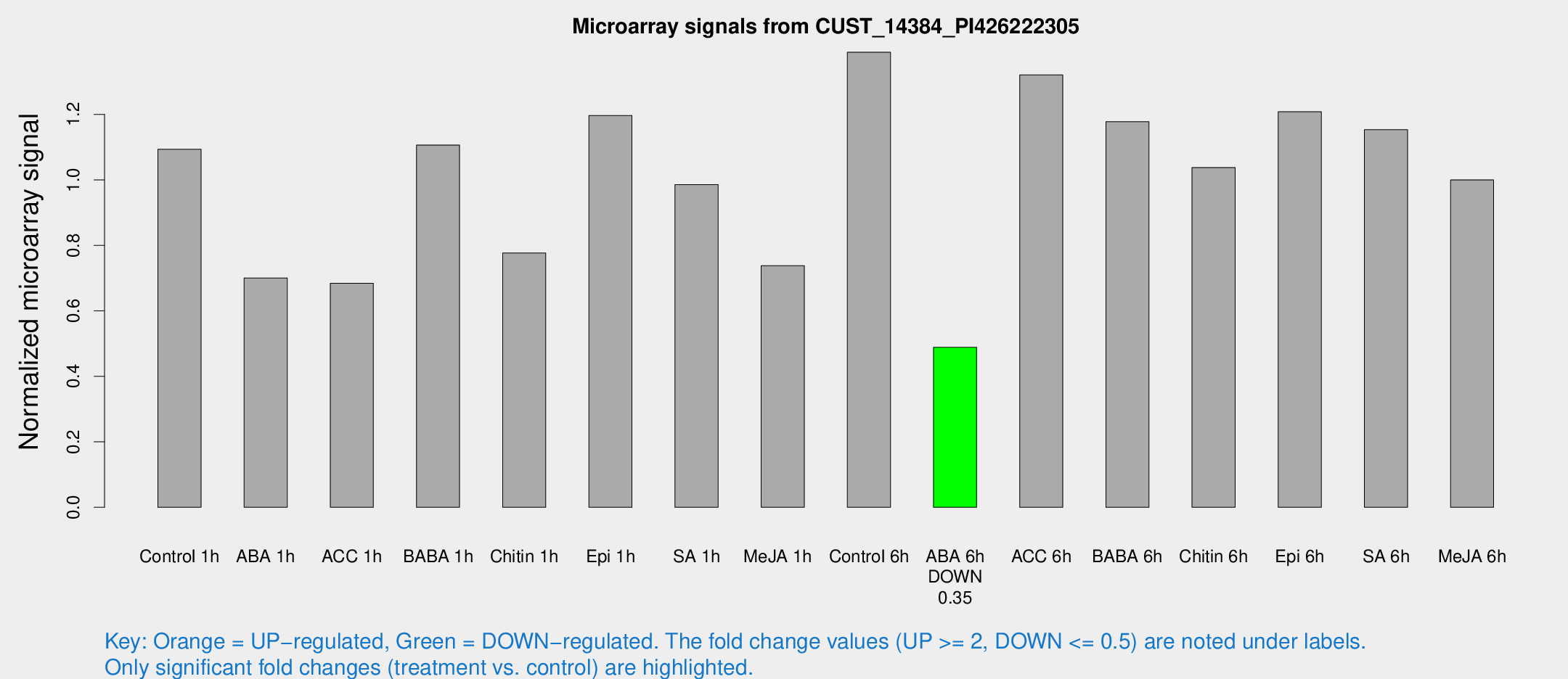
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 811.799 | 89.342 | 1.09344 | 0.063271 |
| ABA 1h | 455.65 | 27.2044 | 0.700302 | 0.04068 |
| ACC 1h | 548.208 | 123.626 | 0.68402 | 0.131095 |
| BABA 1h | 821.017 | 169.521 | 1.10635 | 0.143566 |
| Chitin 1h | 514.125 | 33.2999 | 0.776679 | 0.0724584 |
| Epi 1h | 768.876 | 85.1144 | 1.19622 | 0.129784 |
| SA 1h | 742.51 | 43.017 | 0.985201 | 0.0570222 |
| Me-JA 1h | 446.056 | 45.713 | 0.737974 | 0.0516903 |
| Control 6h | 1090.1 | 285.066 | 1.38978 | 0.290701 |
| ABA 6h | 383.865 | 33.7796 | 0.488544 | 0.0285115 |
| ACC 6h | 1135.25 | 172.362 | 1.32059 | 0.0895833 |
| BABA 6h | 973.036 | 79.1907 | 1.17773 | 0.0681285 |
| Chitin 6h | 813.154 | 55.0175 | 1.03762 | 0.0600685 |
| Epi 6h | 1004.84 | 78.8944 | 1.20808 | 0.0698791 |
| SA 6h | 840.59 | 63.5403 | 1.15308 | 0.107812 |
| Me-JA 6h | 749.506 | 114.742 | 1 | 0.105359 |
Source Transcript PGSC0003DMT400059947 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G53590.1 | +1 | 2e-16 | 72 | 30/66 (45%) | SAUR-like auxin-responsive protein family | chr5:21772107-21772535 FORWARD LENGTH=142 |
