Probe CUST_14334_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14334_PI426222305 | JHI_St_60k_v1 | DMT400060245 | CTGAGTTAAGGAAATCCAGGCTTTGAAAACCTTATATATTACTCGTATATGTATCGCGAA |
All Microarray Probes Designed to Gene DMG400023436
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14379_PI426222305 | JHI_St_60k_v1 | DMT400060244 | CCTGATGAAGTACACTTGGGGACAATCGCGAAAACTGAGTTAAGGAAATCCAGGCTTTGA |
| CUST_14334_PI426222305 | JHI_St_60k_v1 | DMT400060245 | CTGAGTTAAGGAAATCCAGGCTTTGAAAACCTTATATATTACTCGTATATGTATCGCGAA |
Microarray Signals from CUST_14334_PI426222305
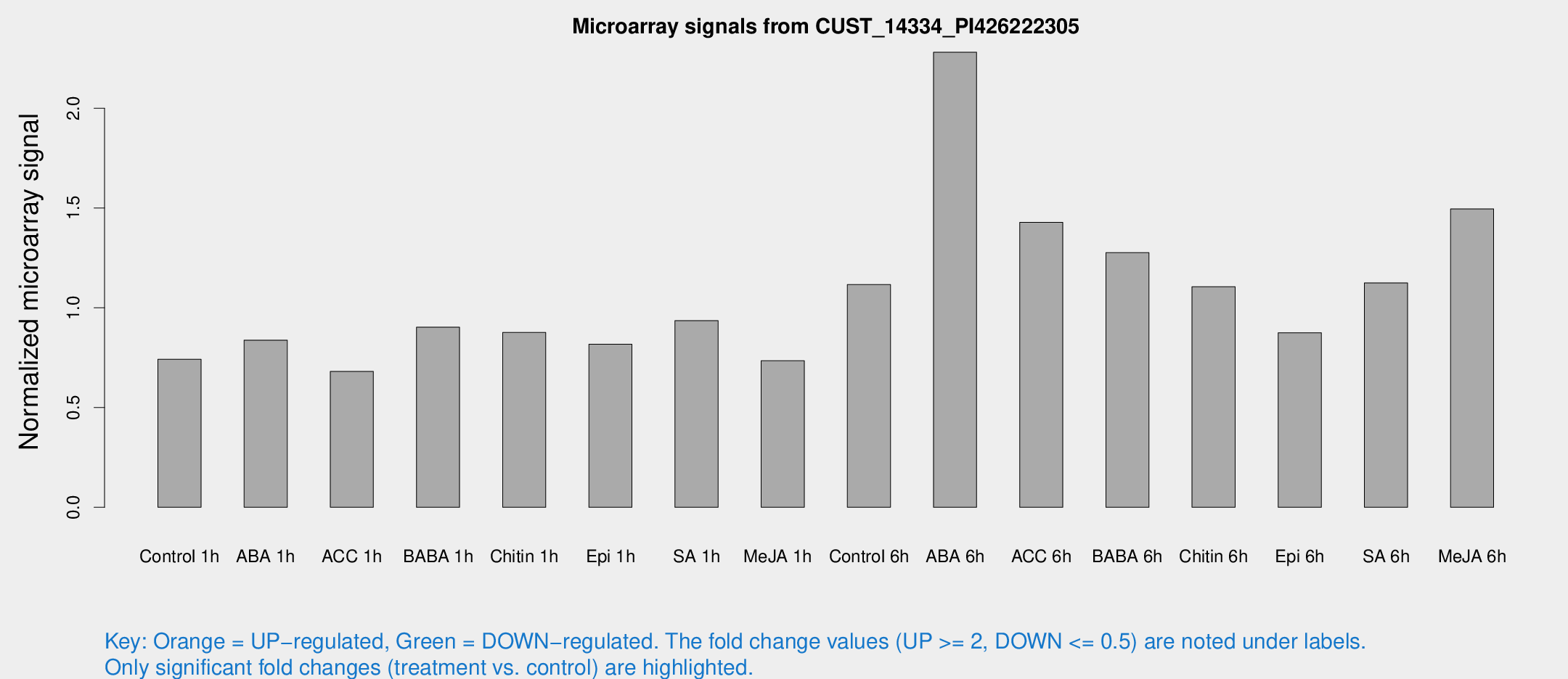
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1371.27 | 134 | 0.742272 | 0.0428906 |
| ABA 1h | 1393.56 | 212.335 | 0.837951 | 0.0980329 |
| ACC 1h | 1353.99 | 294.285 | 0.681164 | 0.120015 |
| BABA 1h | 1701.51 | 388.279 | 0.902804 | 0.151283 |
| Chitin 1h | 1459.64 | 162.879 | 0.876276 | 0.0506276 |
| Epi 1h | 1319.86 | 174.496 | 0.817164 | 0.108619 |
| SA 1h | 1816.77 | 334.42 | 0.93566 | 0.111061 |
| Me-JA 1h | 1121.88 | 162.237 | 0.734464 | 0.0465973 |
| Control 6h | 2176.17 | 482.803 | 1.11619 | 0.191073 |
| ABA 6h | 4526.05 | 618.846 | 2.28103 | 0.195214 |
| ACC 6h | 3037.95 | 341.879 | 1.42831 | 0.0824843 |
| BABA 6h | 2652.27 | 304.567 | 1.27625 | 0.102017 |
| Chitin 6h | 2163.52 | 133.266 | 1.10591 | 0.0650172 |
| Epi 6h | 1829.2 | 203.638 | 0.874917 | 0.0734342 |
| SA 6h | 2069.52 | 254.802 | 1.12474 | 0.0649675 |
| Me-JA 6h | 2888.4 | 631.827 | 1.49562 | 0.257504 |
Source Transcript PGSC0003DMT400060245 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G06810.1 | +3 | 0.0 | 1219 | 590/828 (71%) | acyl-CoA dehydrogenase-related | chr3:2146534-2150654 FORWARD LENGTH=824 |
