Probe CUST_14145_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14145_PI426222305 | JHI_St_60k_v1 | DMT400060273 | CACACGAATACTACGAATTCCATAGGCCTGTCCTTTGGTAACAACACCATCACCTAAGTA |
All Microarray Probes Designed to Gene DMG401023446
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14145_PI426222305 | JHI_St_60k_v1 | DMT400060273 | CACACGAATACTACGAATTCCATAGGCCTGTCCTTTGGTAACAACACCATCACCTAAGTA |
Microarray Signals from CUST_14145_PI426222305
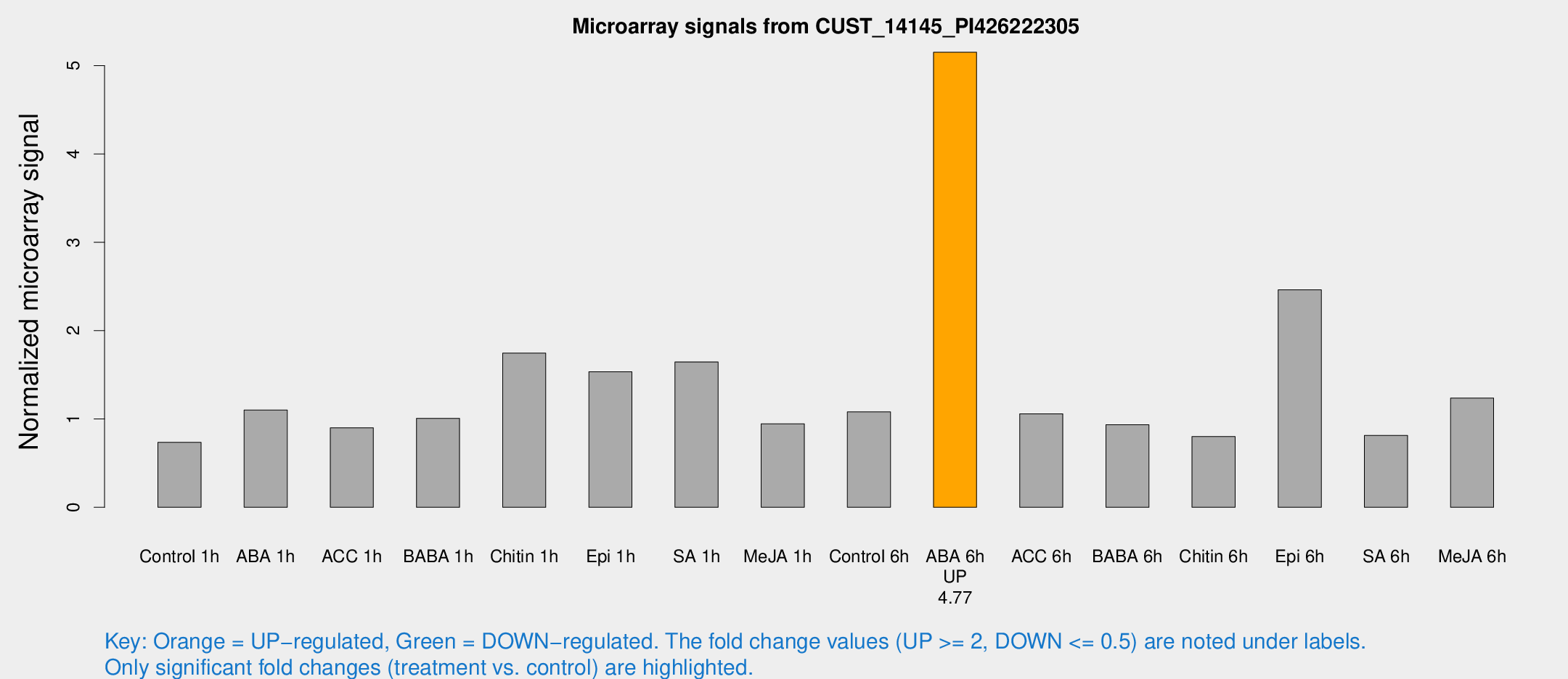
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.38791 | 3.1283 | 0.735422 | 0.426483 |
| ABA 1h | 7.49081 | 3.02458 | 1.1005 | 0.494837 |
| ACC 1h | 6.86795 | 3.47034 | 0.900156 | 0.467932 |
| BABA 1h | 7.18264 | 3.29103 | 1.00666 | 0.475376 |
| Chitin 1h | 12.6225 | 3.44242 | 1.74477 | 0.796282 |
| Epi 1h | 9.73738 | 3.12465 | 1.53388 | 0.495467 |
| SA 1h | 12.6855 | 3.16263 | 1.64542 | 0.436069 |
| Me-JA 1h | 5.69957 | 3.20076 | 0.944526 | 0.531673 |
| Control 6h | 8.75314 | 3.33879 | 1.07945 | 0.508572 |
| ABA 6h | 40.9593 | 6.24785 | 5.15101 | 0.787137 |
| ACC 6h | 9.53712 | 3.99514 | 1.05834 | 0.503134 |
| BABA 6h | 7.82506 | 3.63611 | 0.934833 | 0.457424 |
| Chitin 6h | 6.23915 | 3.61459 | 0.80044 | 0.463474 |
| Epi 6h | 37.0004 | 25.8415 | 2.46259 | 3.13787 |
| SA 6h | 5.89224 | 3.41339 | 0.813829 | 0.47139 |
| Me-JA 6h | 10.1817 | 3.68492 | 1.23608 | 0.508212 |
Source Transcript PGSC0003DMT400060273 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G21400.1 | -2 | 1e-36 | 94 | 44/85 (52%) | Thiamin diphosphate-binding fold (THDP-binding) superfamily protein | chr1:7493492-7496240 FORWARD LENGTH=472 |
