Probe CUST_14059_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14059_PI426222305 | JHI_St_60k_v1 | DMT400059936 | ATAAAACTAATTTTGAAACAGGGGTTCCCAAAACGTTATTCTCAGAAATCTTGCCAGTGT |
All Microarray Probes Designed to Gene DMG400023319
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14059_PI426222305 | JHI_St_60k_v1 | DMT400059936 | ATAAAACTAATTTTGAAACAGGGGTTCCCAAAACGTTATTCTCAGAAATCTTGCCAGTGT |
| CUST_14076_PI426222305 | JHI_St_60k_v1 | DMT400059935 | GAAAATGAATAAAACAGAATGTTTTGGGTGTCTGACATGGGCACATTATGCCTGCAGTCT |
Microarray Signals from CUST_14059_PI426222305
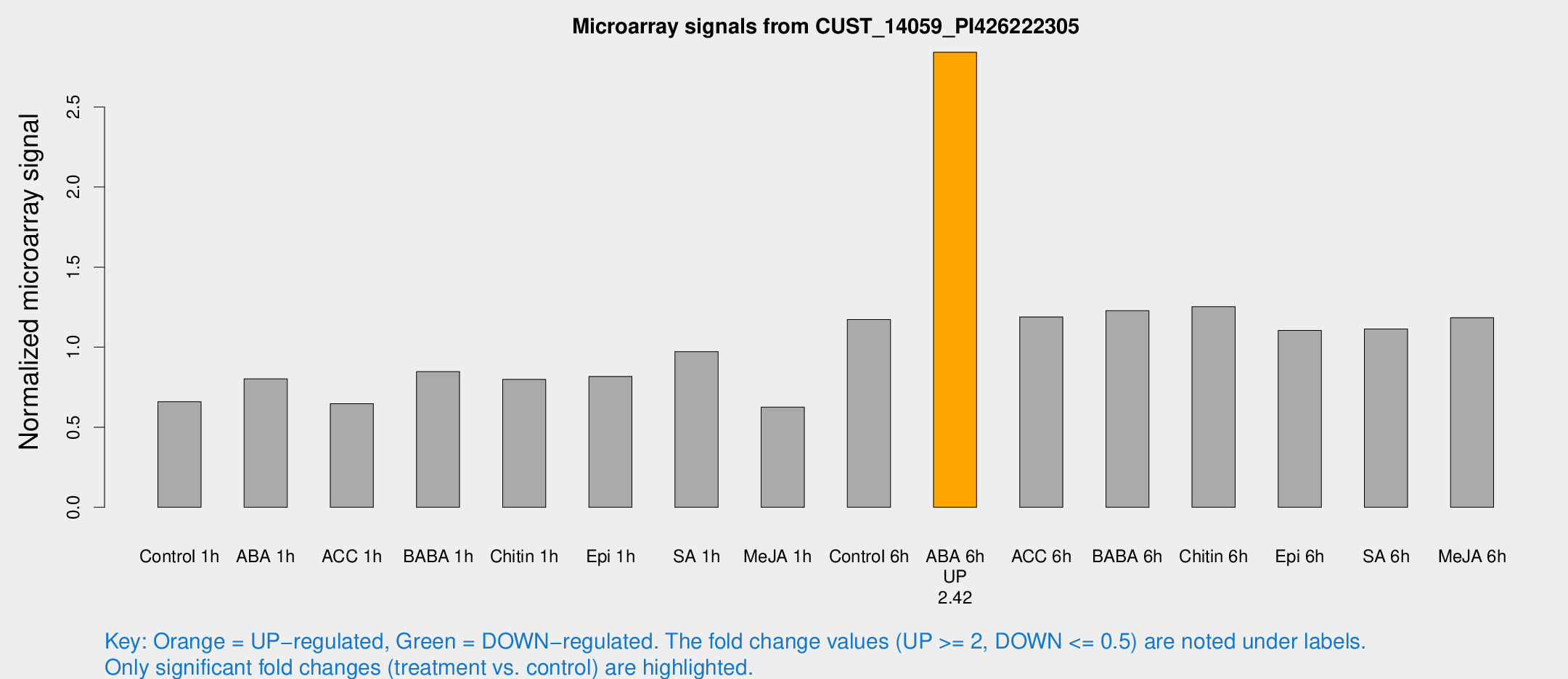
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 49.8267 | 4.2477 | 0.659305 | 0.056547 |
| ABA 1h | 55.6198 | 10.3324 | 0.802588 | 0.122899 |
| ACC 1h | 53.4567 | 12.4952 | 0.647444 | 0.130528 |
| BABA 1h | 65.9953 | 15.5848 | 0.847332 | 0.153478 |
| Chitin 1h | 56.0992 | 10.8016 | 0.79849 | 0.0942462 |
| Epi 1h | 53.715 | 5.36296 | 0.817259 | 0.0847834 |
| SA 1h | 78.183 | 15.0079 | 0.972201 | 0.140446 |
| Me-JA 1h | 38.4301 | 3.89039 | 0.625834 | 0.0633181 |
| Control 6h | 92.5218 | 17.933 | 1.17293 | 0.152345 |
| ABA 6h | 229.274 | 21.1639 | 2.84152 | 0.169273 |
| ACC 6h | 103.387 | 9.51188 | 1.18859 | 0.081535 |
| BABA 6h | 104.086 | 8.18683 | 1.22814 | 0.082578 |
| Chitin 6h | 100.777 | 7.26622 | 1.25238 | 0.0849049 |
| Epi 6h | 97.0039 | 18.0616 | 1.10484 | 0.176644 |
| SA 6h | 85.1371 | 13.6382 | 1.11354 | 0.0831173 |
| Me-JA 6h | 94.7145 | 21.8997 | 1.18399 | 0.227686 |
Source Transcript PGSC0003DMT400059936 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G63620.1 | +3 | 0.0 | 594 | 298/381 (78%) | GroES-like zinc-binding alcohol dehydrogenase family protein | chr5:25466380-25468296 REVERSE LENGTH=427 |
