Probe CUST_1383_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_1383_PI426222305 | JHI_St_60k_v1 | DMT400003477 | TGGTGAAACTACTGAGTTTCTTAAACTTGGTCCAAGTTTCATGAAAAACCAAAGAGCCAG |
All Microarray Probes Designed to Gene DMG400001374
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_1383_PI426222305 | JHI_St_60k_v1 | DMT400003477 | TGGTGAAACTACTGAGTTTCTTAAACTTGGTCCAAGTTTCATGAAAAACCAAAGAGCCAG |
Microarray Signals from CUST_1383_PI426222305
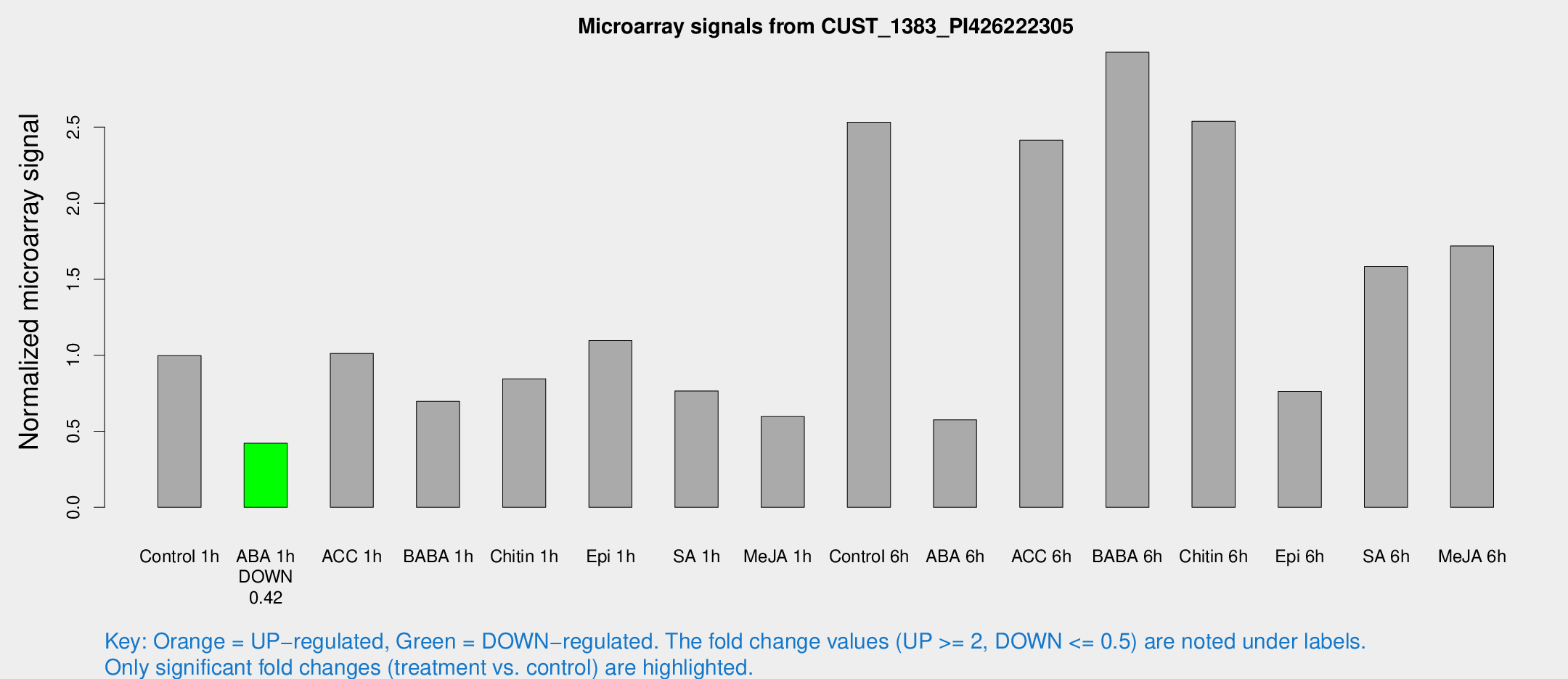
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.5272 | 3.30967 | 0.99795 | 0.214118 |
| ABA 1h | 5.82506 | 3.10601 | 0.421337 | 0.226297 |
| ACC 1h | 18.0985 | 5.86854 | 1.01158 | 0.336927 |
| BABA 1h | 11.2438 | 3.44694 | 0.696839 | 0.290305 |
| Chitin 1h | 12.645 | 3.43848 | 0.845179 | 0.291698 |
| Epi 1h | 15.3252 | 3.26224 | 1.096 | 0.252664 |
| SA 1h | 14.1259 | 4.71607 | 0.765334 | 0.311146 |
| Me-JA 1h | 8.11258 | 3.3415 | 0.596283 | 0.285971 |
| Control 6h | 49.2745 | 18.5929 | 2.53314 | 1.2559 |
| ABA 6h | 10.7009 | 3.82692 | 0.575671 | 0.234669 |
| ACC 6h | 67.093 | 33.1567 | 2.41406 | 2.75879 |
| BABA 6h | 60.3224 | 20.8177 | 2.99308 | 1.25609 |
| Chitin 6h | 42.7184 | 5.86132 | 2.5392 | 0.299684 |
| Epi 6h | 14.64 | 4.30544 | 0.763056 | 0.329709 |
| SA 6h | 30.2069 | 12.2982 | 1.58362 | 1.07409 |
| Me-JA 6h | 31.8525 | 11.6296 | 1.71877 | 0.719601 |
Source Transcript PGSC0003DMT400003477 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc02g089160.2 | +1 | 0.0 | 905 | 439/453 (97%) | genomic_reference:SL2.50ch02 gene_region:45622003-45624825 transcript_region:SL2.50ch02:45622003..45624825+ go_terms:GO:0004497 functional_description:Cytochrome P450 |
| TAIR PP10 | AT5G38970.1 | +1 | 0.0 | 660 | 312/448 (70%) | brassinosteroid-6-oxidase 1 | chr5:15594935-15597774 REVERSE LENGTH=465 |
