Probe CUST_13664_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13664_PI426222305 | JHI_St_60k_v1 | DMT400018046 | CATGTTATGTTGCTTCACTCTTCAAAAATATCGACAGTTGCATCTCAGATCTTTCAAAAG |
All Microarray Probes Designed to Gene DMG400007010
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13664_PI426222305 | JHI_St_60k_v1 | DMT400018046 | CATGTTATGTTGCTTCACTCTTCAAAAATATCGACAGTTGCATCTCAGATCTTTCAAAAG |
Microarray Signals from CUST_13664_PI426222305
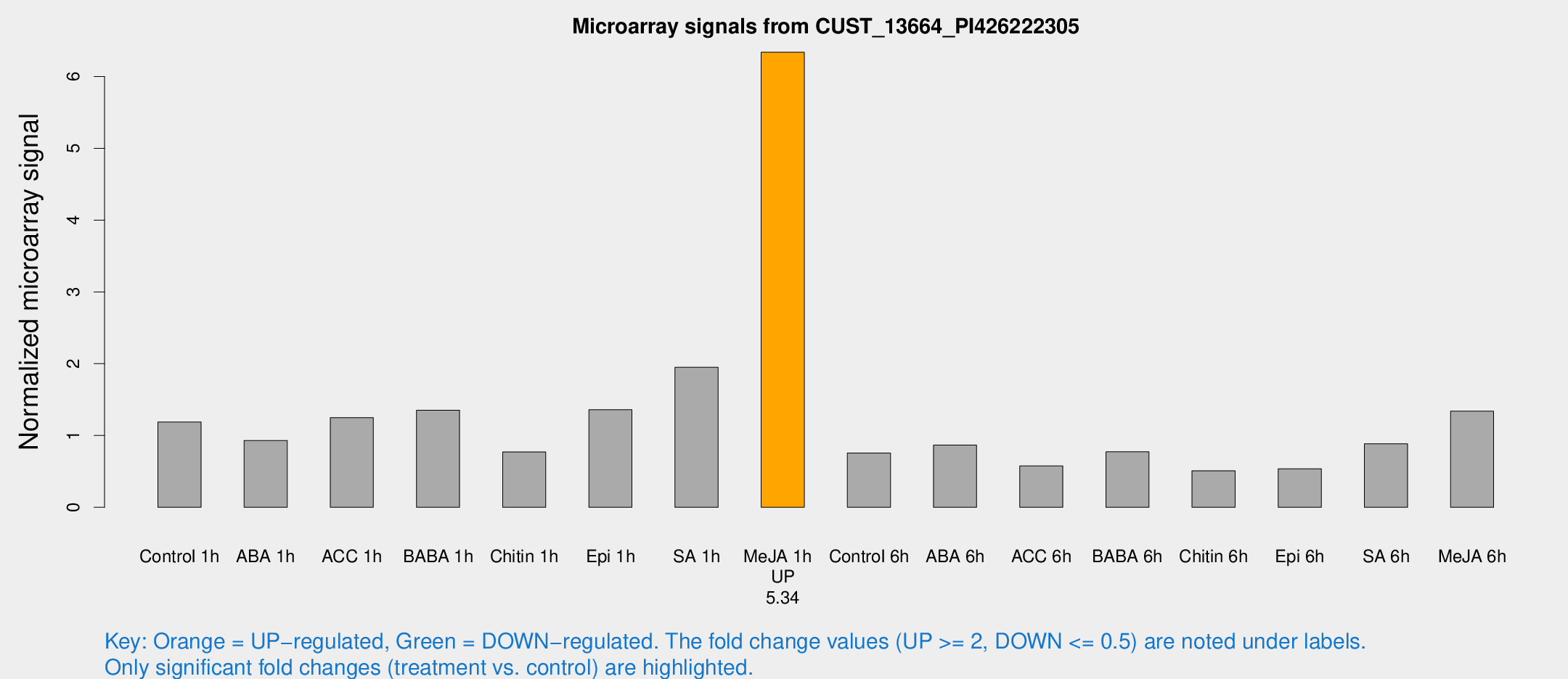
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 54.5157 | 20.4448 | 1.1877 | 0.383135 |
| ABA 1h | 37.0861 | 10.9601 | 0.929875 | 0.286598 |
| ACC 1h | 55.7881 | 15.6234 | 1.24903 | 0.290852 |
| BABA 1h | 54.4534 | 10.6715 | 1.35245 | 0.206431 |
| Chitin 1h | 28.0395 | 3.84884 | 0.770101 | 0.105425 |
| Epi 1h | 47.4484 | 4.34097 | 1.35863 | 0.124745 |
| SA 1h | 81.0323 | 5.80545 | 1.95006 | 0.221042 |
| Me-JA 1h | 209.637 | 14.6424 | 6.33799 | 0.381275 |
| Control 6h | 31.426 | 5.1708 | 0.754896 | 0.107414 |
| ABA 6h | 38.4586 | 7.4834 | 0.866335 | 0.115527 |
| ACC 6h | 28.9497 | 7.26397 | 0.576174 | 0.180392 |
| BABA 6h | 37.362 | 8.66646 | 0.773053 | 0.238865 |
| Chitin 6h | 22.0298 | 4.20383 | 0.508052 | 0.0997158 |
| Epi 6h | 26.038 | 6.79409 | 0.536837 | 0.128048 |
| SA 6h | 38.517 | 11.5453 | 0.885607 | 0.197737 |
| Me-JA 6h | 54.9991 | 8.66158 | 1.33855 | 0.125634 |
Source Transcript PGSC0003DMT400018046 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G01260.1 | +2 | 2e-176 | 526 | 307/601 (51%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr1:109595-111367 FORWARD LENGTH=590 |
