Probe CUST_13202_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13202_PI426222305 | JHI_St_60k_v1 | DMT400006324 | TAATGTGGGCAAAGCCGCTGGTATCCCTAGAGTTTGTGGCGTAAACATTCCTTACAAGAT |
All Microarray Probes Designed to Gene DMG400002471
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13202_PI426222305 | JHI_St_60k_v1 | DMT400006324 | TAATGTGGGCAAAGCCGCTGGTATCCCTAGAGTTTGTGGCGTAAACATTCCTTACAAGAT |
Microarray Signals from CUST_13202_PI426222305
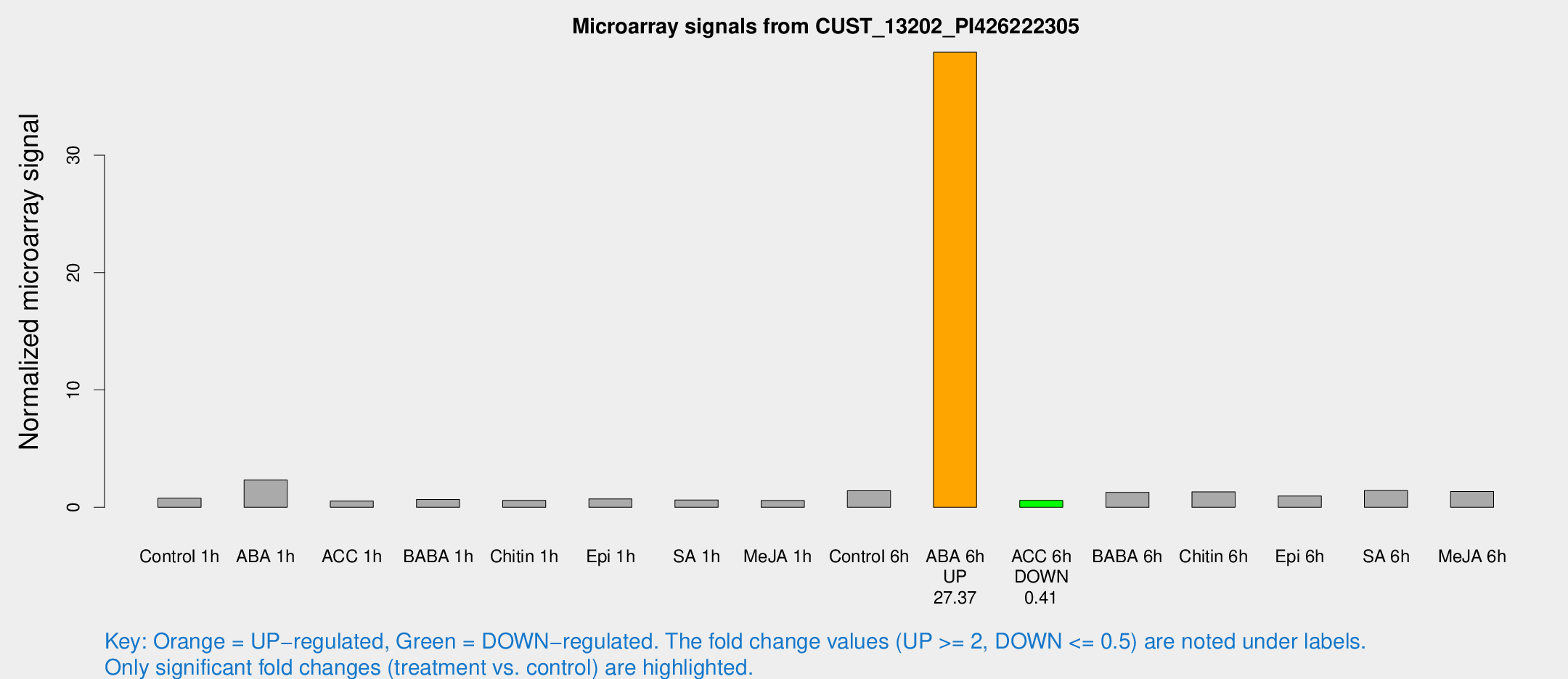
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 329.731 | 19.3809 | 0.772814 | 0.0853612 |
| ABA 1h | 931.79 | 224.585 | 2.3212 | 0.549405 |
| ACC 1h | 281.284 | 100.307 | 0.524872 | 0.257134 |
| BABA 1h | 277.379 | 28.7156 | 0.668882 | 0.108413 |
| Chitin 1h | 248.99 | 79.6892 | 0.58502 | 0.184873 |
| Epi 1h | 307.969 | 98.9811 | 0.71596 | 0.342365 |
| SA 1h | 273.085 | 16.1352 | 0.623802 | 0.052073 |
| Me-JA 1h | 207.534 | 42.1764 | 0.575276 | 0.0702735 |
| Control 6h | 640.981 | 158.409 | 1.41678 | 0.237679 |
| ABA 6h | 18393.2 | 3699.1 | 38.7829 | 7.58703 |
| ACC 6h | 293.407 | 46.5968 | 0.58641 | 0.142817 |
| BABA 6h | 614.649 | 60.9724 | 1.27722 | 0.165672 |
| Chitin 6h | 613.026 | 95.8607 | 1.31835 | 0.197587 |
| Epi 6h | 465.627 | 46.236 | 0.960782 | 0.101866 |
| SA 6h | 619.462 | 113.533 | 1.4205 | 0.132444 |
| Me-JA 6h | 592.131 | 105.837 | 1.35544 | 0.194525 |
Source Transcript PGSC0003DMT400006324 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G38540.1 | +2 | 1e-24 | 96 | 59/113 (52%) | lipid transfer protein 1 | chr2:16130418-16130893 FORWARD LENGTH=118 |
