Probe CUST_13192_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13192_PI426222305 | JHI_St_60k_v1 | DMT400007591 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
All Microarray Probes Designed to Gene DMG400002930
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13157_PI426222305 | JHI_St_60k_v1 | DMT400007592 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
| CUST_13192_PI426222305 | JHI_St_60k_v1 | DMT400007591 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
Microarray Signals from CUST_13192_PI426222305
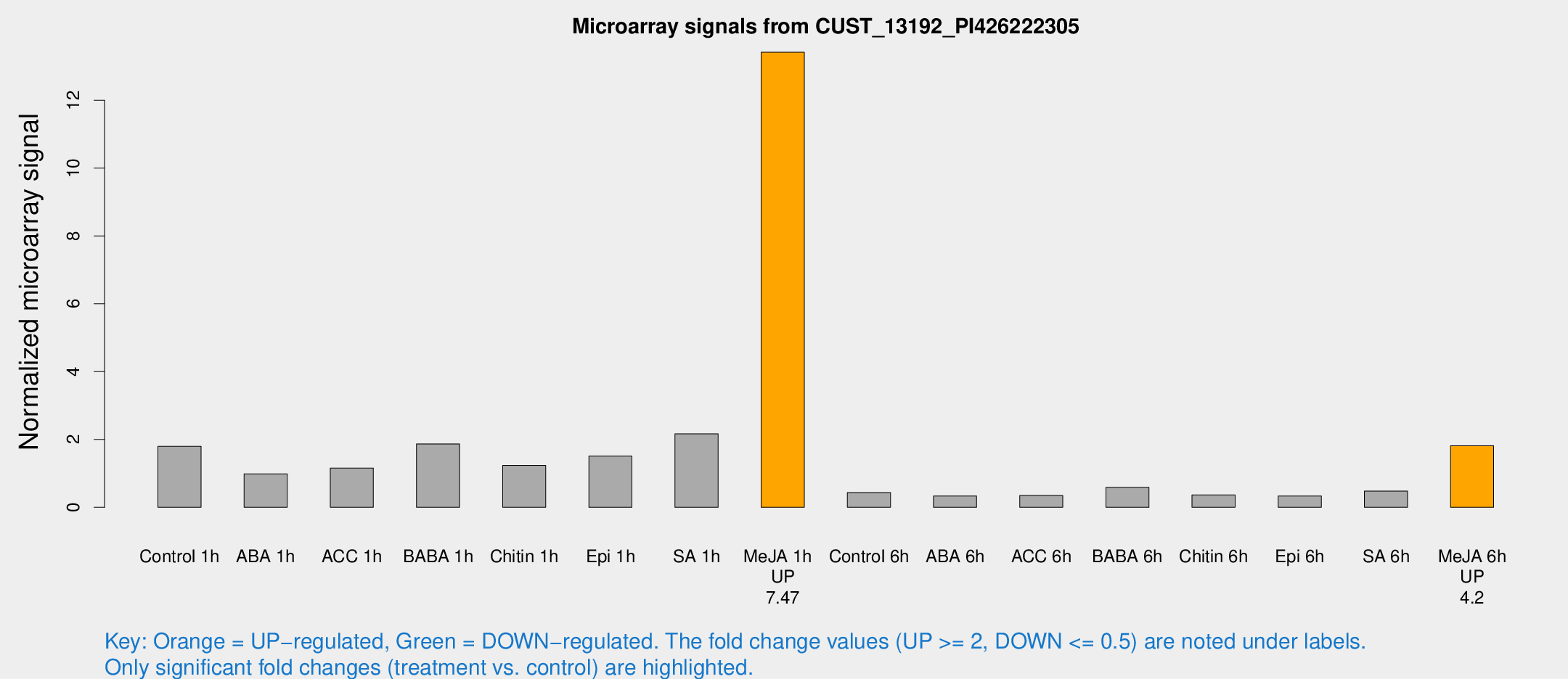
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2323.35 | 755.446 | 1.79708 | 0.457935 |
| ABA 1h | 1190.67 | 438.131 | 0.98507 | 0.35262 |
| ACC 1h | 1567.45 | 516.685 | 1.15508 | 0.353231 |
| BABA 1h | 2215.16 | 428.809 | 1.86923 | 0.216834 |
| Chitin 1h | 1326.87 | 168.65 | 1.23452 | 0.153867 |
| Epi 1h | 1562.35 | 196.481 | 1.50963 | 0.173836 |
| SA 1h | 2627.26 | 182.898 | 2.16551 | 0.151017 |
| Me-JA 1h | 13294.6 | 2387.76 | 13.4171 | 1.24206 |
| Control 6h | 577.135 | 184.439 | 0.4326 | 0.126475 |
| ABA 6h | 439.221 | 102.486 | 0.33452 | 0.0577896 |
| ACC 6h | 481.227 | 83.2836 | 0.347065 | 0.0946642 |
| BABA 6h | 827.476 | 216.466 | 0.587519 | 0.152282 |
| Chitin 6h | 457.129 | 36.2236 | 0.363043 | 0.0373293 |
| Epi 6h | 441.952 | 26.904 | 0.33222 | 0.0194586 |
| SA 6h | 563.445 | 62.8615 | 0.47863 | 0.0289203 |
| Me-JA 6h | 2128.21 | 123.047 | 1.81645 | 0.189027 |
Source Transcript PGSC0003DMT400007591 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19180.1 | +2 | 1e-23 | 99 | 71/173 (41%) | jasmonate-zim-domain protein 1 | chr1:6622312-6623271 FORWARD LENGTH=253 |
