Probe CUST_13157_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13157_PI426222305 | JHI_St_60k_v1 | DMT400007592 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
All Microarray Probes Designed to Gene DMG400002930
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13157_PI426222305 | JHI_St_60k_v1 | DMT400007592 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
| CUST_13192_PI426222305 | JHI_St_60k_v1 | DMT400007591 | GACCTGAAGATTATAGGCCTATTGCTTTGATATACTACTATTAGATGACCAAGTTTTTGG |
Microarray Signals from CUST_13157_PI426222305
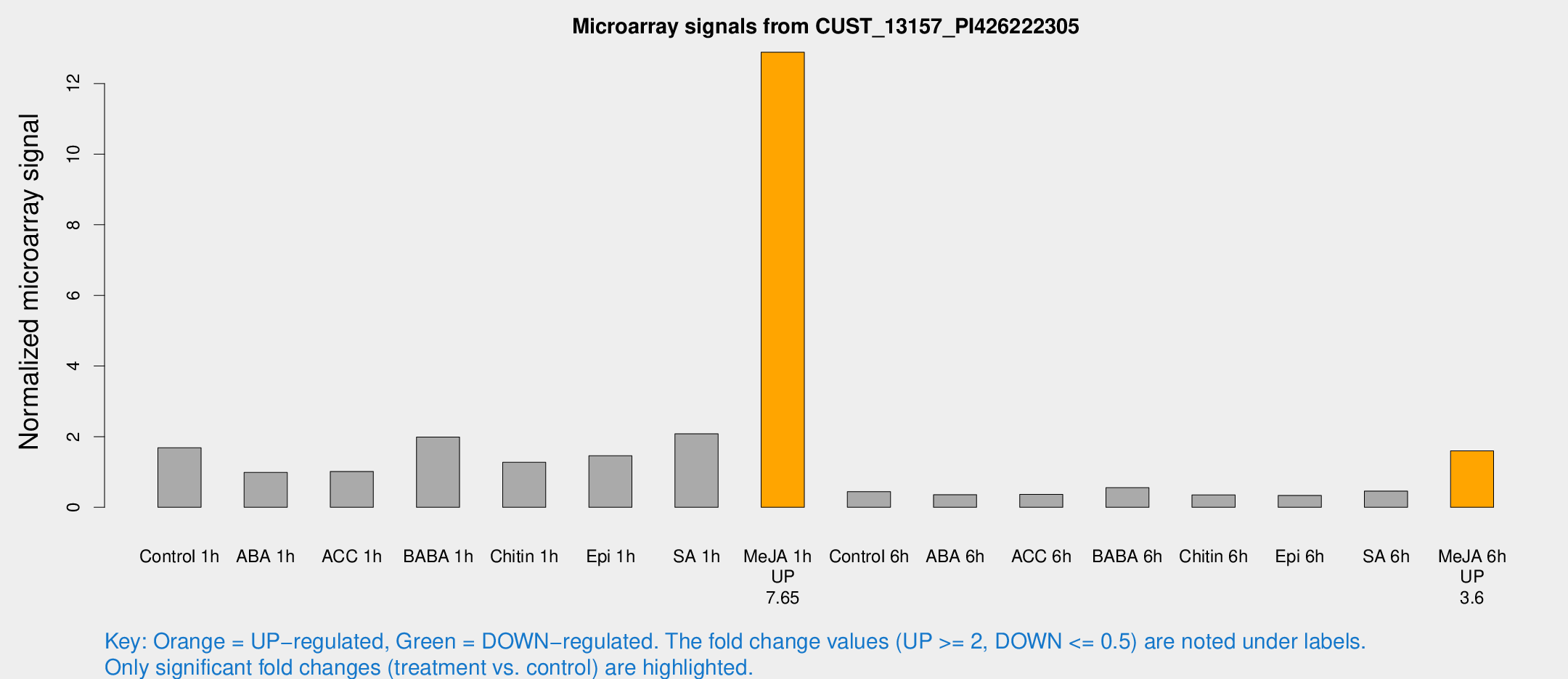
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2316.98 | 767.472 | 1.6837 | 0.451202 |
| ABA 1h | 1223.91 | 400.353 | 0.986944 | 0.297398 |
| ACC 1h | 1493.5 | 529.971 | 1.01512 | 0.352588 |
| BABA 1h | 2548.69 | 584.454 | 1.98786 | 0.334022 |
| Chitin 1h | 1432.1 | 82.9694 | 1.27508 | 0.0736661 |
| Epi 1h | 1592.05 | 174.078 | 1.45782 | 0.143039 |
| SA 1h | 2678.67 | 209.062 | 2.08098 | 0.160079 |
| Me-JA 1h | 13565.9 | 2471.58 | 12.8862 | 1.31935 |
| Control 6h | 632.69 | 205.431 | 0.443572 | 0.137867 |
| ABA 6h | 498.829 | 120.694 | 0.35691 | 0.0635299 |
| ACC 6h | 542.421 | 108.962 | 0.365484 | 0.107411 |
| BABA 6h | 825.423 | 206.164 | 0.55423 | 0.146954 |
| Chitin 6h | 467.224 | 30.0838 | 0.350973 | 0.0269482 |
| Epi 6h | 477.306 | 47.9345 | 0.336517 | 0.0308947 |
| SA 6h | 576.478 | 72.9635 | 0.460386 | 0.0277537 |
| Me-JA 6h | 1990.4 | 134.194 | 1.59705 | 0.189935 |
Source Transcript PGSC0003DMT400007592 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19180.1 | +1 | 2e-33 | 127 | 105/264 (40%) | jasmonate-zim-domain protein 1 | chr1:6622312-6623271 FORWARD LENGTH=253 |
