Probe CUST_12949_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12949_PI426222305 | JHI_St_60k_v1 | DMT400063327 | CTGGATTATTCATGAGCAAACATAATCATGCTACATTATTCCTTCTTCACAGGTACAACA |
All Microarray Probes Designed to Gene DMG400024632
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12890_PI426222305 | JHI_St_60k_v1 | DMT400063326 | GATCAGCCTAATTCGAATTTGTCTTTAGAGCTTGGAGAAGGAAGAGATAAGATGCCCCTA |
| CUST_12949_PI426222305 | JHI_St_60k_v1 | DMT400063327 | CTGGATTATTCATGAGCAAACATAATCATGCTACATTATTCCTTCTTCACAGGTACAACA |
Microarray Signals from CUST_12949_PI426222305
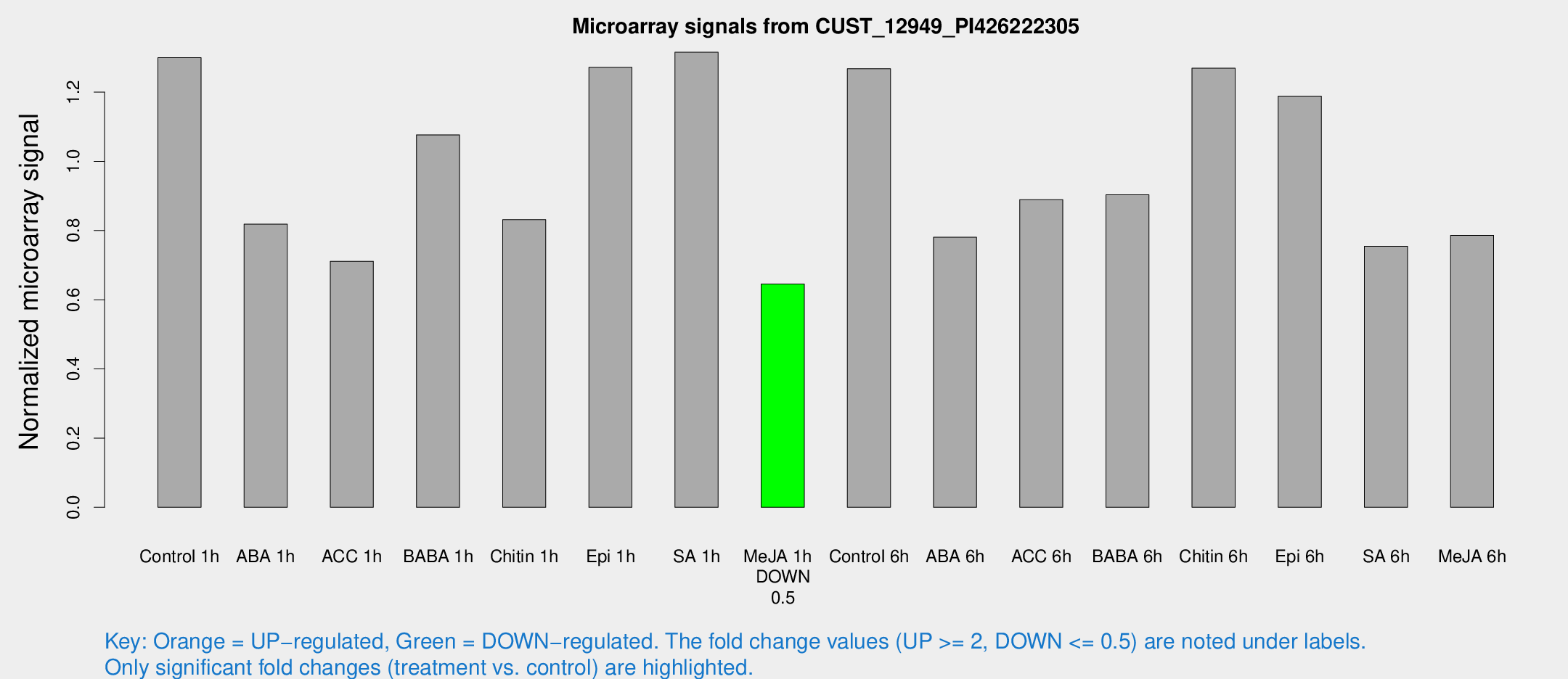
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 105.278 | 11.1574 | 1.2999 | 0.0863408 |
| ABA 1h | 59.2304 | 8.37428 | 0.818296 | 0.123233 |
| ACC 1h | 76.0829 | 32.4293 | 0.71089 | 0.415865 |
| BABA 1h | 87.4883 | 17.9762 | 1.07659 | 0.143903 |
| Chitin 1h | 60.6071 | 6.51204 | 0.831545 | 0.12551 |
| Epi 1h | 89.3548 | 10.108 | 1.27174 | 0.133557 |
| SA 1h | 109.176 | 10.656 | 1.31543 | 0.0865094 |
| Me-JA 1h | 43.0732 | 6.06334 | 0.645587 | 0.065784 |
| Control 6h | 104.407 | 15.9633 | 1.2679 | 0.105293 |
| ABA 6h | 66.8656 | 5.35706 | 0.780646 | 0.0611646 |
| ACC 6h | 82.8202 | 9.98926 | 0.889419 | 0.0678853 |
| BABA 6h | 83.5614 | 13.8005 | 0.903398 | 0.134889 |
| Chitin 6h | 110.499 | 15.3626 | 1.26917 | 0.164821 |
| Epi 6h | 112.978 | 26.7477 | 1.18852 | 0.299441 |
| SA 6h | 60.5462 | 6.68128 | 0.75459 | 0.0644475 |
| Me-JA 6h | 68.7301 | 18.2123 | 0.78608 | 0.199677 |
Source Transcript PGSC0003DMT400063327 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G02150.2 | +2 | 8e-45 | 162 | 107/271 (39%) | plastid transcription factor 1 | chr3:391522-392589 FORWARD LENGTH=355 |
