Probe CUST_12796_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12796_PI426222305 | JHI_St_60k_v1 | DMT400063116 | CCCCGCTTTTCTACCTCAAGTAAGCAATTTATGCTTGTTTCAAATTAGAGTTTAACTTCT |
All Microarray Probes Designed to Gene DMG400024554
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12931_PI426222305 | JHI_St_60k_v1 | DMT400063117 | CTACACAGATAATAGGCAGCAAGTTGAACCAGGGAGACCAGACAGAATAGAAACTGAGTC |
| CUST_12796_PI426222305 | JHI_St_60k_v1 | DMT400063116 | CCCCGCTTTTCTACCTCAAGTAAGCAATTTATGCTTGTTTCAAATTAGAGTTTAACTTCT |
Microarray Signals from CUST_12796_PI426222305
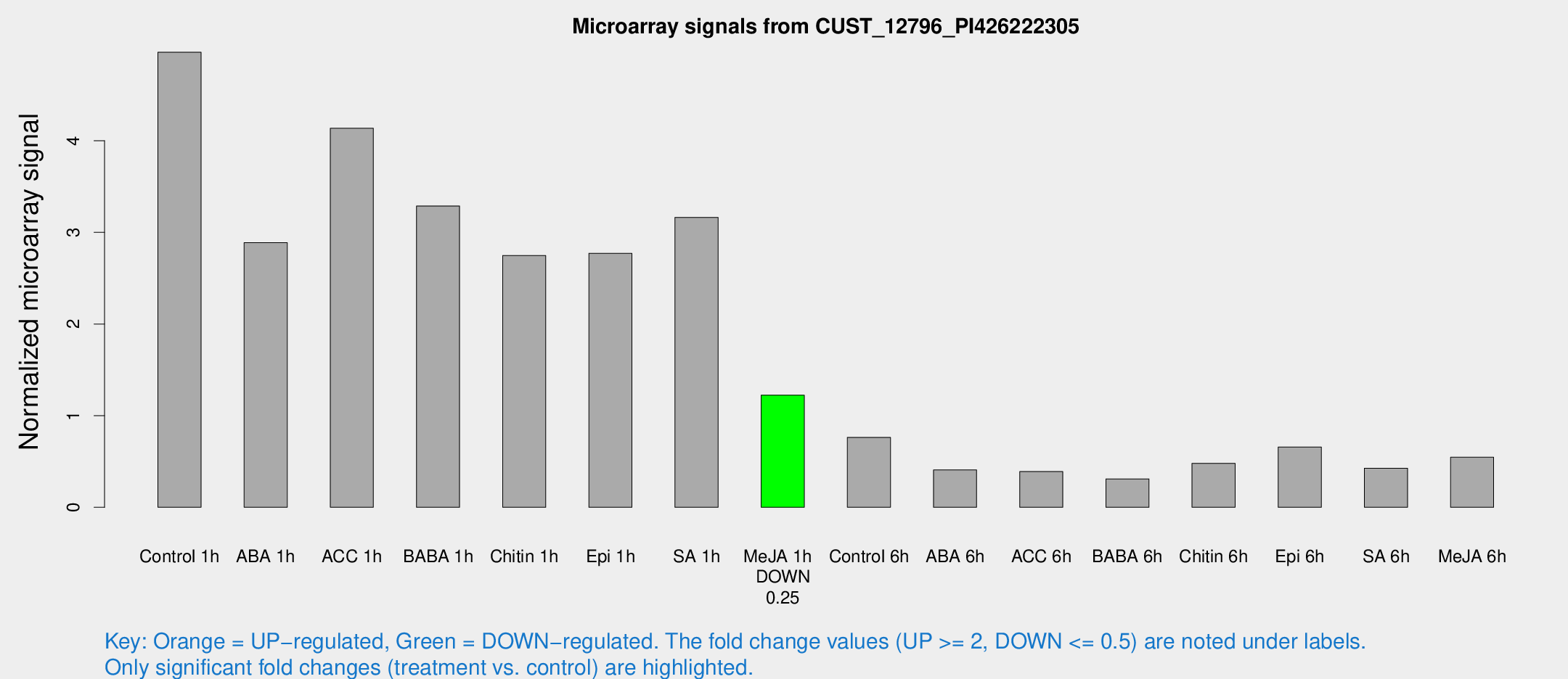
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 97.3 | 6.8445 | 4.96555 | 0.322316 |
| ABA 1h | 50.3225 | 5.20714 | 2.88687 | 0.454781 |
| ACC 1h | 83.2115 | 5.77521 | 4.1357 | 0.566082 |
| BABA 1h | 62.5964 | 6.76665 | 3.28802 | 0.250676 |
| Chitin 1h | 48.5815 | 4.27253 | 2.7476 | 0.227834 |
| Epi 1h | 48.7125 | 9.19224 | 2.77155 | 0.563933 |
| SA 1h | 63.5361 | 4.67457 | 3.16323 | 0.327395 |
| Me-JA 1h | 20.2348 | 3.6721 | 1.22421 | 0.24946 |
| Control 6h | 15.807 | 3.94123 | 0.763464 | 0.171149 |
| ABA 6h | 8.87748 | 3.14163 | 0.408279 | 0.162527 |
| ACC 6h | 8.90415 | 3.67308 | 0.390418 | 0.167883 |
| BABA 6h | 6.85379 | 3.37885 | 0.309852 | 0.157235 |
| Chitin 6h | 10.6797 | 3.363 | 0.480188 | 0.181449 |
| Epi 6h | 14.6572 | 3.5505 | 0.657709 | 0.169341 |
| SA 6h | 8.53039 | 3.19603 | 0.42707 | 0.175681 |
| Me-JA 6h | 12.1182 | 4.02713 | 0.546149 | 0.190066 |
Source Transcript PGSC0003DMT400063116 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G00050.1 | +3 | 7e-31 | 124 | 112/274 (41%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr4:17863-19848 FORWARD LENGTH=399 |
