Probe CUST_12735_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12735_PI426222305 | JHI_St_60k_v1 | DMT400063103 | GAGGCGAATGTTATTAGTTTTTCGCCGTTTTTATTTTTGTTTCCTCTGTATCAGTCCCAT |
All Microarray Probes Designed to Gene DMG400024547
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12812_PI426222305 | JHI_St_60k_v1 | DMT400063104 | ATGACGAGTTATGAATTTTTGACTTGATCTCTAGGATGCAGAAGGTGTTCTTGTAGCCAA |
| CUST_12735_PI426222305 | JHI_St_60k_v1 | DMT400063103 | GAGGCGAATGTTATTAGTTTTTCGCCGTTTTTATTTTTGTTTCCTCTGTATCAGTCCCAT |
Microarray Signals from CUST_12735_PI426222305
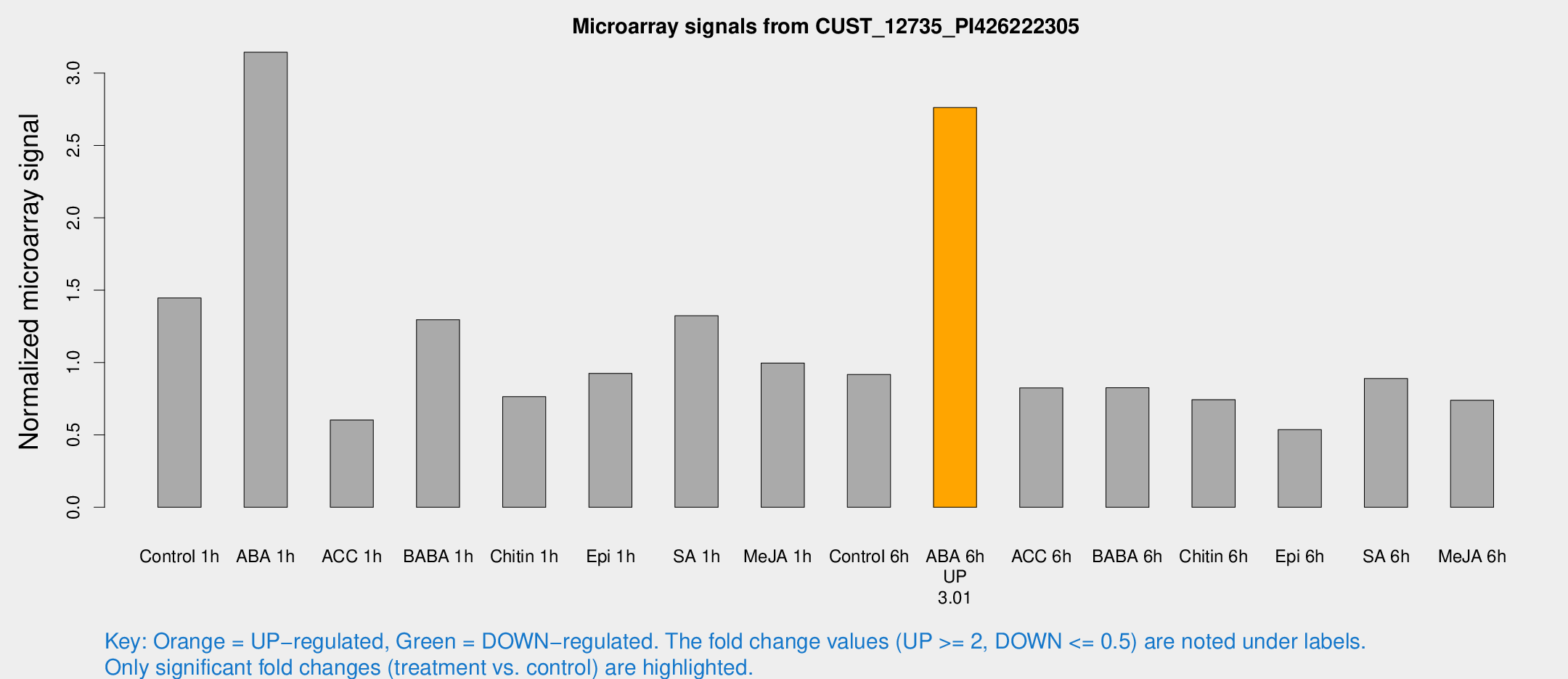
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 46.5316 | 8.4162 | 1.44656 | 0.187504 |
| ABA 1h | 91.1352 | 18.9104 | 3.14366 | 0.615573 |
| ACC 1h | 21.3474 | 5.76767 | 0.603246 | 0.194045 |
| BABA 1h | 41.5893 | 10.9313 | 1.29617 | 0.249235 |
| Chitin 1h | 23.7599 | 6.55772 | 0.764511 | 0.30436 |
| Epi 1h | 25.4161 | 3.64447 | 0.925155 | 0.140753 |
| SA 1h | 42.5685 | 4.23093 | 1.32313 | 0.132006 |
| Me-JA 1h | 25.6775 | 3.76332 | 0.996423 | 0.146851 |
| Control 6h | 31.1387 | 9.38489 | 0.917522 | 0.249042 |
| ABA 6h | 98.8812 | 24.7708 | 2.76162 | 0.686433 |
| ACC 6h | 31.5831 | 8.49137 | 0.824438 | 0.32427 |
| BABA 6h | 30.6588 | 7.94074 | 0.825654 | 0.197944 |
| Chitin 6h | 24.7937 | 4.13853 | 0.74379 | 0.125251 |
| Epi 6h | 19.0557 | 4.31764 | 0.536768 | 0.120861 |
| SA 6h | 28.9453 | 6.55659 | 0.889355 | 0.144886 |
| Me-JA 6h | 26.0844 | 7.87458 | 0.739841 | 0.239307 |
Source Transcript PGSC0003DMT400063103 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G07920.1 | +1 | 0.0 | 1035 | 541/737 (73%) | diacylglycerol kinase1 | chr5:2525197-2528396 REVERSE LENGTH=728 |
