Probe CUST_12474_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12474_PI426222305 | JHI_St_60k_v1 | DMT400003003 | AAAGTTGGTTTTTCCATTCCATGTTGGCTGAGAAAATGAGTTAAAATGGTGATTAGCGTG |
All Microarray Probes Designed to Gene DMG400001182
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12305_PI426222305 | JHI_St_60k_v1 | DMT400003001 | ACAAGTCCCGTCTTTCACTTTGGAGCCTTCCAAAAGGAATACCCAAGTTGGTTAGATGCA |
| CUST_12474_PI426222305 | JHI_St_60k_v1 | DMT400003003 | AAAGTTGGTTTTTCCATTCCATGTTGGCTGAGAAAATGAGTTAAAATGGTGATTAGCGTG |
Microarray Signals from CUST_12474_PI426222305
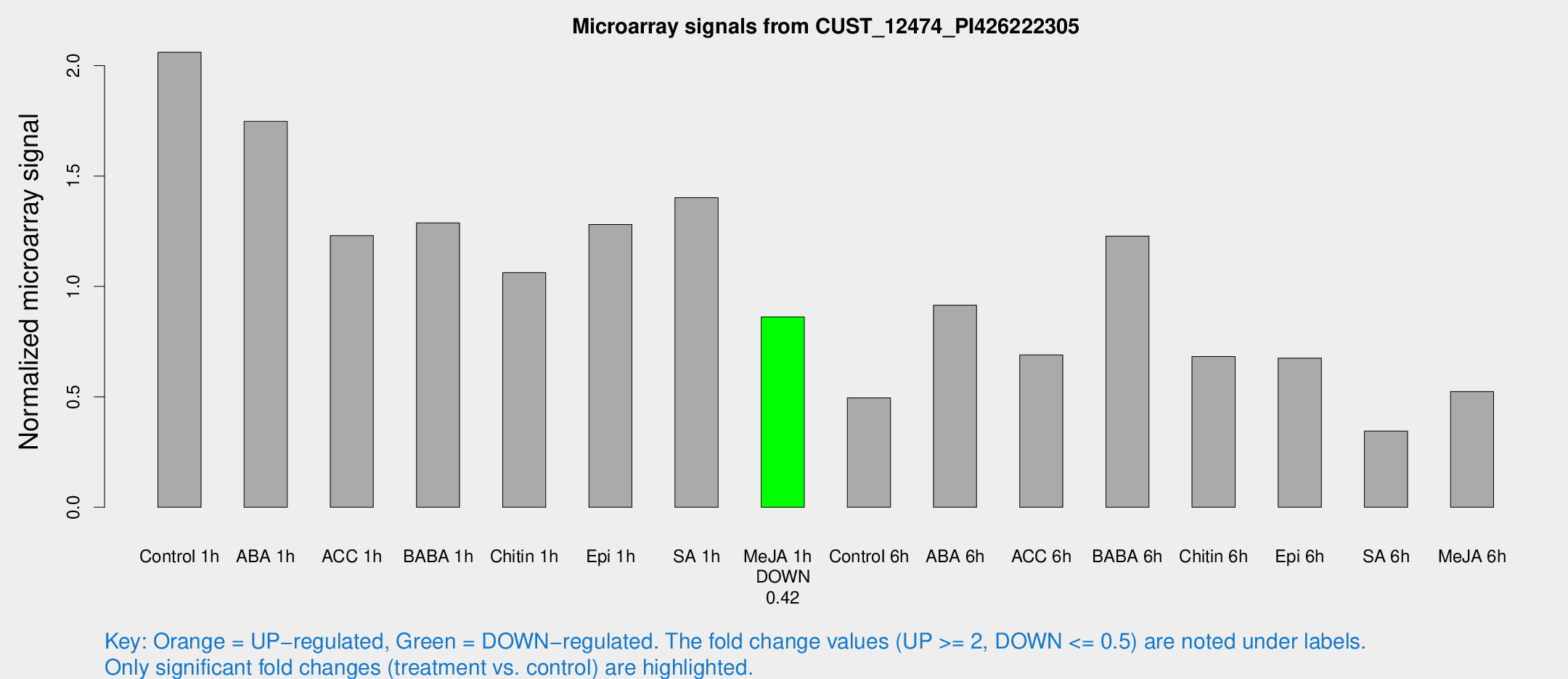
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 393.766 | 35.8737 | 2.06044 | 0.120322 |
| ABA 1h | 299.707 | 46.7846 | 1.7475 | 0.19858 |
| ACC 1h | 267.296 | 75.3252 | 1.22996 | 0.365402 |
| BABA 1h | 243.915 | 43.1284 | 1.288 | 0.119887 |
| Chitin 1h | 181.494 | 11.0386 | 1.06291 | 0.0644537 |
| Epi 1h | 210.52 | 12.6255 | 1.28078 | 0.0766777 |
| SA 1h | 274.278 | 20.5712 | 1.40198 | 0.0828838 |
| Me-JA 1h | 136.528 | 22.1304 | 0.861684 | 0.0733377 |
| Control 6h | 97.1184 | 16.5495 | 0.495498 | 0.0440537 |
| ABA 6h | 187.551 | 25.2199 | 0.915368 | 0.0697533 |
| ACC 6h | 151.242 | 13.8541 | 0.689881 | 0.0797643 |
| BABA 6h | 264.846 | 34.4419 | 1.22822 | 0.112418 |
| Chitin 6h | 137.939 | 8.89374 | 0.682629 | 0.044001 |
| Epi 6h | 144.785 | 9.38829 | 0.675276 | 0.0557455 |
| SA 6h | 71.0744 | 19.0388 | 0.344841 | 0.0748118 |
| Me-JA 6h | 104.419 | 23.9622 | 0.524491 | 0.0795411 |
Source Transcript PGSC0003DMT400003003 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19310.1 | +3 | 7e-68 | 218 | 105/193 (54%) | RING/U-box superfamily protein | chr1:6676424-6677104 REVERSE LENGTH=226 |
