Probe CUST_12421_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12421_PI426222305 | JHI_St_60k_v1 | DMT400063715 | GATAGCCGAAGCTCTTAATCCGAGCTCGCTATTTTTTGTTCTTTTCTTTTGGTGTGTTTT |
All Microarray Probes Designed to Gene DMG400024764
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12438_PI426222305 | JHI_St_60k_v1 | DMT400063716 | GCCGAAGCTCTTAATCCGAGCTCGCTATTTTTTGTTCTTTTCTTTTGGTGTGTTTTCTAA |
| CUST_12377_PI426222305 | JHI_St_60k_v1 | DMT400063717 | GCCGAAGCTCTTAATCCGAGCTCGCTATTTTTTGTTCTTTTCTTTTGGTGTGTTTTCTAA |
| CUST_12421_PI426222305 | JHI_St_60k_v1 | DMT400063715 | GATAGCCGAAGCTCTTAATCCGAGCTCGCTATTTTTTGTTCTTTTCTTTTGGTGTGTTTT |
Microarray Signals from CUST_12421_PI426222305
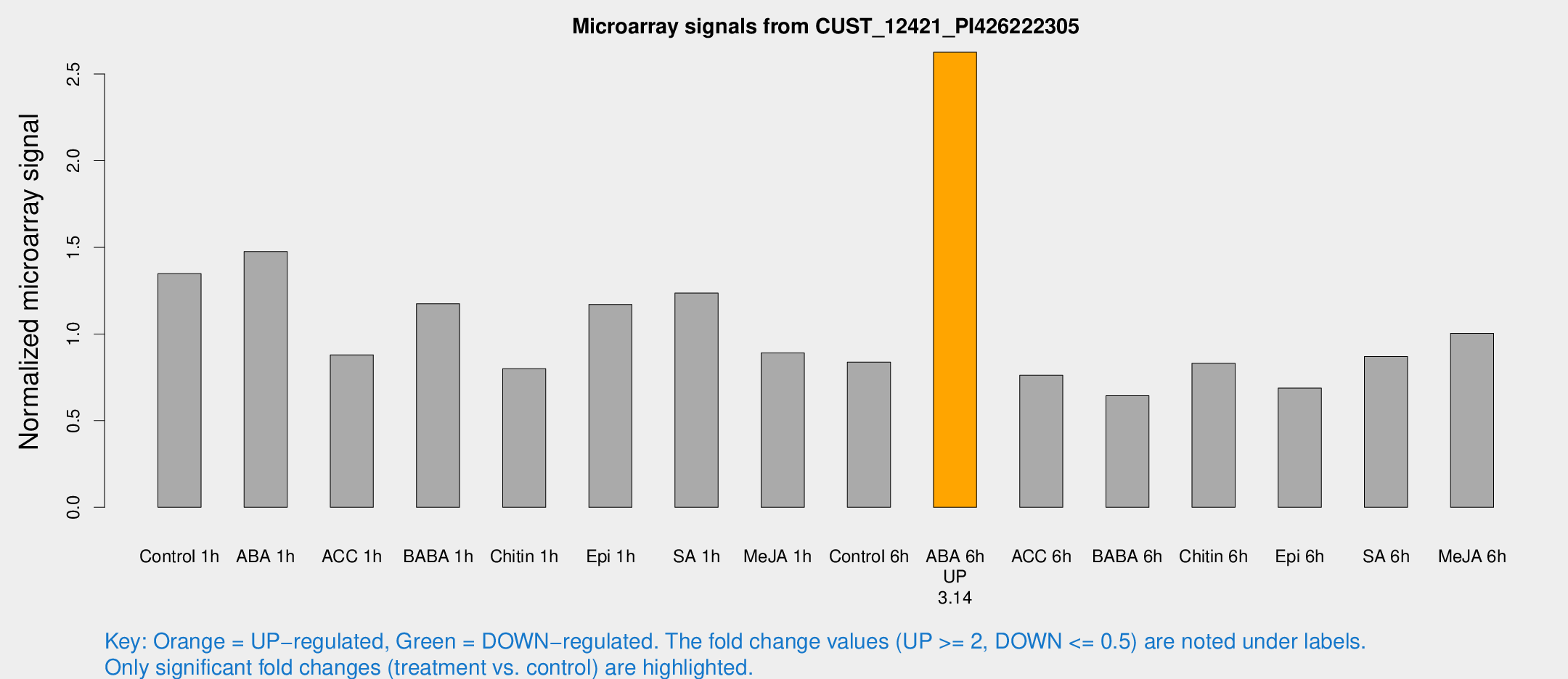
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 197.398 | 27.1429 | 1.34778 | 0.0980997 |
| ABA 1h | 190.331 | 21.0036 | 1.47557 | 0.0979793 |
| ACC 1h | 146.13 | 44.04 | 0.87888 | 0.267636 |
| BABA 1h | 166.171 | 22.397 | 1.17454 | 0.0733724 |
| Chitin 1h | 103.492 | 6.93307 | 0.799429 | 0.0534624 |
| Epi 1h | 146.381 | 9.87638 | 1.1701 | 0.0729171 |
| SA 1h | 183.867 | 15.2654 | 1.23566 | 0.10835 |
| Me-JA 1h | 105.732 | 10.6567 | 0.890211 | 0.122516 |
| Control 6h | 122.443 | 13.9887 | 0.837309 | 0.0552891 |
| ABA 6h | 405.614 | 38.7956 | 2.62543 | 0.169905 |
| ACC 6h | 125.935 | 8.47954 | 0.761547 | 0.123729 |
| BABA 6h | 107.275 | 20.5874 | 0.643472 | 0.111334 |
| Chitin 6h | 127.843 | 8.50943 | 0.830807 | 0.0834236 |
| Epi 6h | 112.354 | 9.74421 | 0.687431 | 0.0579957 |
| SA 6h | 124.404 | 8.30824 | 0.870142 | 0.161981 |
| Me-JA 6h | 147.12 | 21.6168 | 1.00414 | 0.14921 |
Source Transcript PGSC0003DMT400063715 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
