Probe CUST_12374_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12374_PI426222305 | JHI_St_60k_v1 | DMT400063769 | CTATCAAGCTAATGCTGTCGCCTTCACCACAACGGCATCAAGATAGTGAGTAATAATTAT |
All Microarray Probes Designed to Gene DMG400024781
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12374_PI426222305 | JHI_St_60k_v1 | DMT400063769 | CTATCAAGCTAATGCTGTCGCCTTCACCACAACGGCATCAAGATAGTGAGTAATAATTAT |
Microarray Signals from CUST_12374_PI426222305
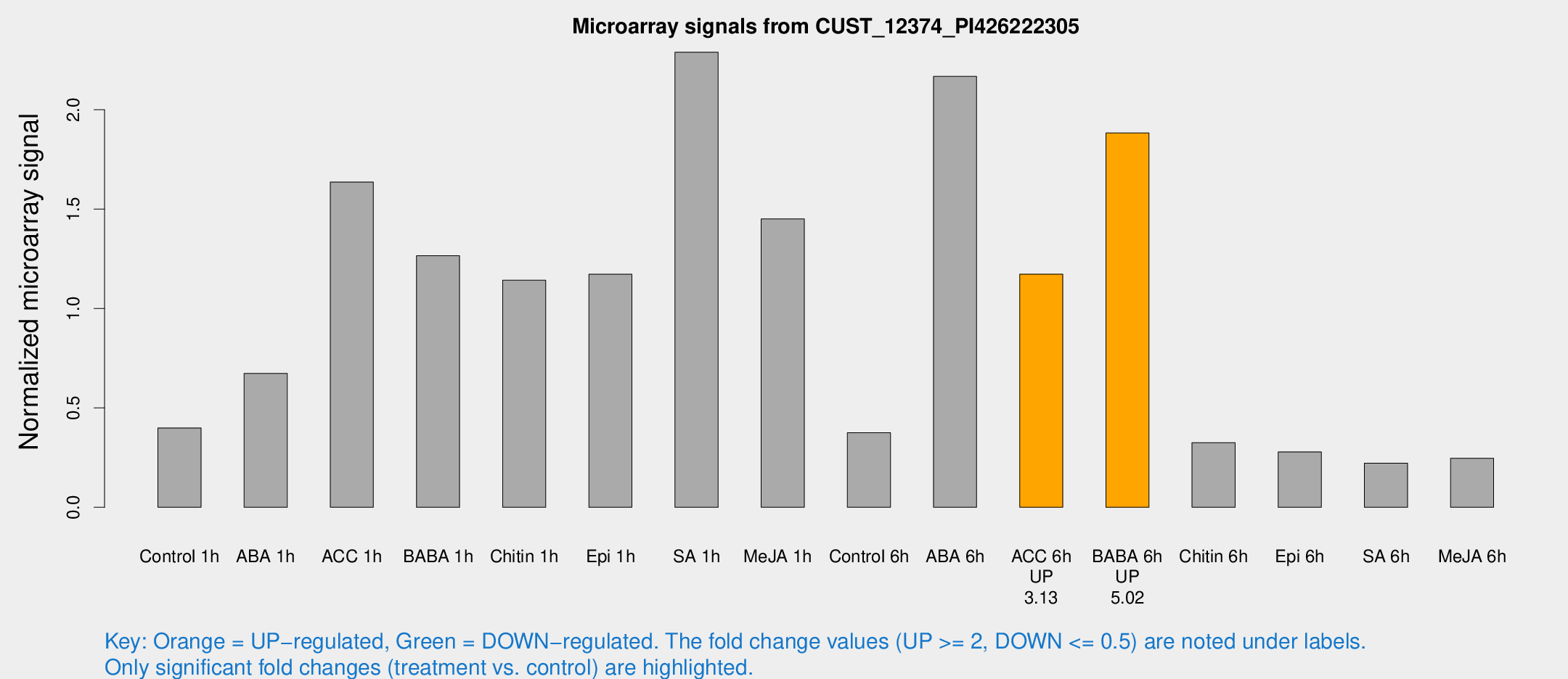
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 16.5289 | 5.76037 | 0.399021 | 0.213213 |
| ABA 1h | 22.0608 | 5.26202 | 0.673237 | 0.140919 |
| ACC 1h | 62.9611 | 15.4251 | 1.63653 | 0.36782 |
| BABA 1h | 44.1324 | 9.49629 | 1.26532 | 0.191623 |
| Chitin 1h | 35.7895 | 3.69507 | 1.14258 | 0.154565 |
| Epi 1h | 35.4921 | 3.7638 | 1.17226 | 0.127827 |
| SA 1h | 86.595 | 20.2351 | 2.28944 | 0.450135 |
| Me-JA 1h | 41.248 | 3.96921 | 1.4511 | 0.140441 |
| Control 6h | 13.4516 | 3.2283 | 0.375232 | 0.0982741 |
| ABA 6h | 96.9039 | 33.8625 | 2.16782 | 1.11001 |
| ACC 6h | 47.052 | 4.75681 | 1.17305 | 0.148111 |
| BABA 6h | 81.3996 | 28.1418 | 1.88306 | 0.589458 |
| Chitin 6h | 12.7773 | 3.57184 | 0.325161 | 0.107915 |
| Epi 6h | 12.6273 | 5.09876 | 0.278877 | 0.115286 |
| SA 6h | 8.15535 | 3.36423 | 0.221942 | 0.106477 |
| Me-JA 6h | 9.05127 | 3.1979 | 0.246981 | 0.0989822 |
Source Transcript PGSC0003DMT400063769 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G29060.1 | +3 | 3e-174 | 523 | 289/647 (45%) | GRAS family transcription factor | chr2:12481991-12484075 FORWARD LENGTH=694 |
