Probe CUST_12320_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12320_PI426222305 | JHI_St_60k_v1 | DMT400063719 | TTTATATGTTTCTTATACCGTACCGTTGAATTTTGGGCAGGAACTGGGAGCTGACAGCCC |
All Microarray Probes Designed to Gene DMG400024765
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12617_PI426222305 | JHI_St_60k_v1 | DMT400063718 | TTTGACCAAAAGGGATTCACTGGTGGACTAAACTACTACCGCGCTCTTGATTTGAACTGG |
| CUST_12320_PI426222305 | JHI_St_60k_v1 | DMT400063719 | TTTATATGTTTCTTATACCGTACCGTTGAATTTTGGGCAGGAACTGGGAGCTGACAGCCC |
Microarray Signals from CUST_12320_PI426222305
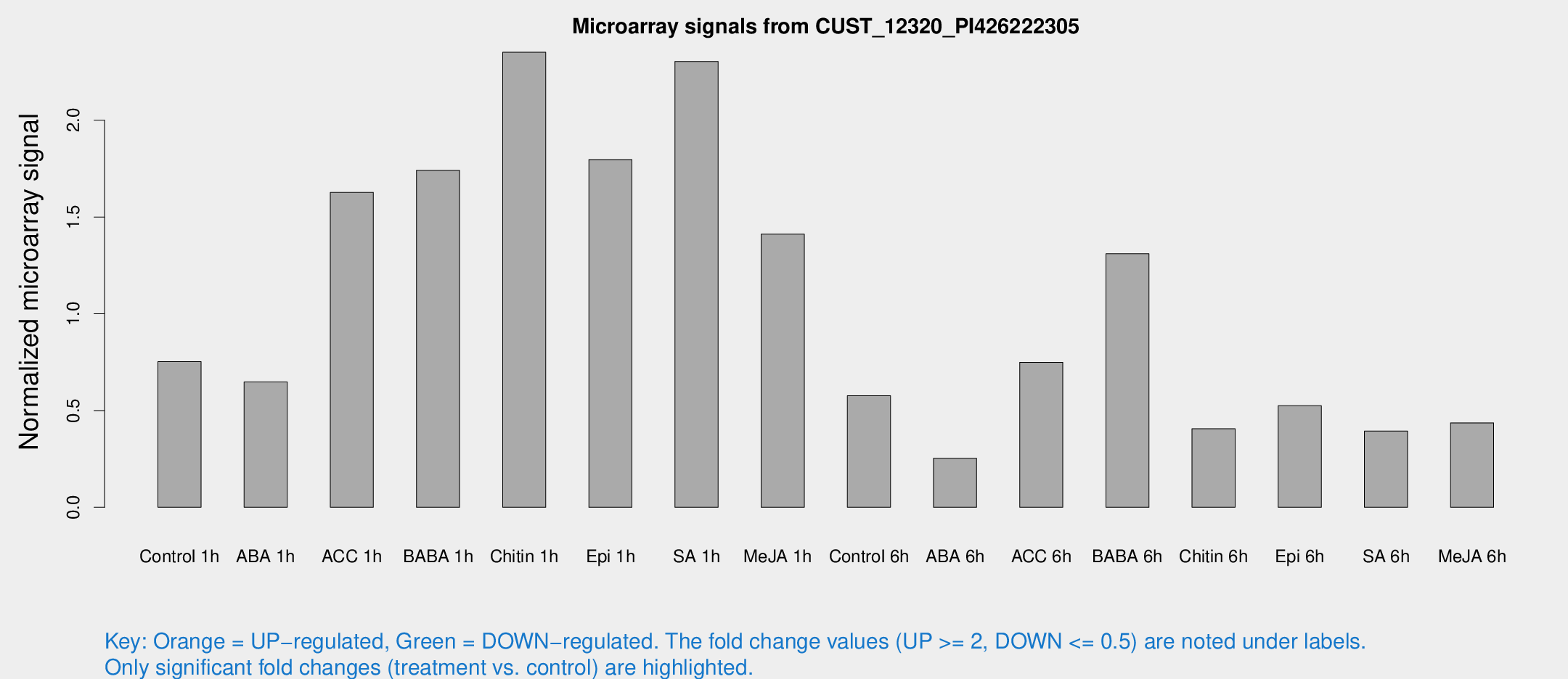
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.8536 | 5.70553 | 0.752582 | 0.126772 |
| ABA 1h | 24.97 | 12.3787 | 0.648002 | 0.503796 |
| ACC 1h | 55.3552 | 12.5901 | 1.62696 | 0.380156 |
| BABA 1h | 56.3254 | 13.8751 | 1.74115 | 0.298981 |
| Chitin 1h | 67.5208 | 8.07864 | 2.35155 | 0.426804 |
| Epi 1h | 50.0258 | 7.21804 | 1.79676 | 0.279957 |
| SA 1h | 74.7164 | 5.5789 | 2.30391 | 0.209025 |
| Me-JA 1h | 36.5276 | 4.26686 | 1.41193 | 0.164382 |
| Control 6h | 22.9926 | 9.74523 | 0.576398 | 0.282322 |
| ABA 6h | 8.89051 | 3.95375 | 0.253673 | 0.124492 |
| ACC 6h | 28.4487 | 5.80071 | 0.74943 | 0.161573 |
| BABA 6h | 50.1618 | 15.0919 | 1.31011 | 0.338979 |
| Chitin 6h | 14.4315 | 4.33034 | 0.405818 | 0.145054 |
| Epi 6h | 19.4389 | 4.49168 | 0.524826 | 0.143534 |
| SA 6h | 14.0942 | 5.5498 | 0.393577 | 0.152105 |
| Me-JA 6h | 16.0875 | 6.06974 | 0.43638 | 0.157423 |
Source Transcript PGSC0003DMT400063719 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G05600.1 | +3 | 3e-110 | 306 | 145/243 (60%) | alpha/beta-Hydrolases superfamily protein | chr3:1623485-1624704 REVERSE LENGTH=331 |
